Digital PCR, along with other new techniques such as next-generation sequencing, is transforming microbial research and infectious disease management. The combination of speed, increased sensitivity, and absolute quantitation of microorganisms is advancing many fields, including pathogen identification and detection, public health, epidemiology, phylogeny, and characterization of the composition, and changes in microbiomes.
This page gives a brief overview of notable applications of digital PCR in microbiology, showcasing its broad potential for research and improving public health.
Page Contents
The ability to concurrently detect low concentrations of pathogens, detect variants, profile populations, and provide absolute quantitation is opening up new possibilities for human and animal testing and treatment in both research and clinical settings. Detection and quantification of pathogens and drug resistance are two applications that will benefit from the sensitivity of Droplet Digital™ PCR (ddPCR™) Technology.
Detection and Quantification of Pathogens
Viral load — the ability to detect rare events and obtain absolute quantitation in the absence of a standard curve increases the accuracy of results, virtually eliminates current problems with standardization, and facilitates effective treatment decisions. For example, digital PCR provides more sensitive testing of viral load in patients with HIV/AIDS being treated with combination antiretroviral therapy. Viremia in many of these patients is below the detection limits of commercial assays.
- Studies comparing ddPCR technology with qPCR for both proviral and episomal HIV DNA targets indicate that the ddPCR system yields improved accuracy and precision to well below the limit of quantification by qPCR (Strain et al. 2013). Furthermore, ddPCR technology has been shown to be able to detect a single copy of plasma HIV-1 RNA (Anderson et al. 2012).
- Recently, the first verified cure of a child with HIV was reported (Persaud et al. 2013). After conventional tests detected no virus, the ddPCR System provided a more sensitive assay to confirm that the child was functionally cured and there was no latent viral reservoir.
- The only adult known to have been cured of HIV/AIDS received bone marrow transplants from a donor who had two copies of a natural mutation that inactivates CCR5, an HIV receptor on CD4+ T cells. Sangamo BioSciences, Inc. used its proprietary zinc finger DNA-binding protein (ZFP) technology to engineer gene-editing ZFP nucleases that inactivate CCR5 in HIV patients' own T cells and then infused these altered HIV-resistant cells back into the patients. Bio-Rad's QX100™ Droplet Digital PCR System was then used to determine the proviral HIV DNA levels in patients in Phase 1 and 2 clinical trials of this new therapy.
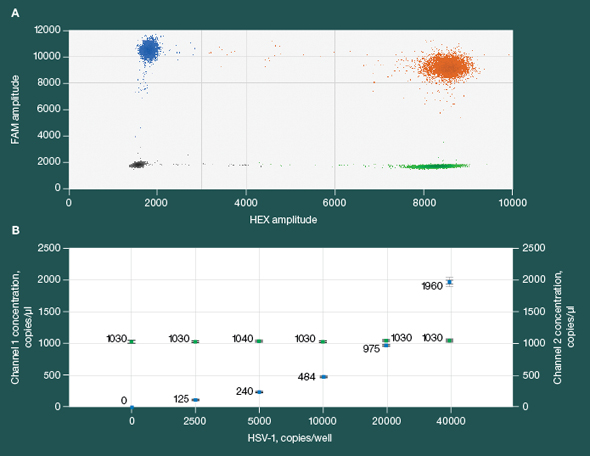
Precise and reproducible detection of herpes simplex virus 1 (HSV-1) and ß2 microglobulin (B2M) targets in a duplex assay A, 2-D fluorescence amplitude plot shows three merged replicate samples of 10,000 copies/well of HSV-1 duplexed with 20,000 copies/well of B2M. The black cluster on the plot represents the negative droplets, the green cluster represents the droplets that are positive for HSV-1 only, the blue cluster represents the droplets that are positive for B2M only, and the orange cluster represents the droplets that are positive for both HSV-1 and B2M targets. B, concentration plot shows merged triplicate wells across an HSV-1 sample dilution series from 0 to 40,000 copies in a constant background of 20,000 copies (66 ng human genomic DNA) of B2M. The blue markers indicate HSV-1 copies/μl and the green markers indicate B2M copies/μl. All error bars are generated by QuantaSoft™ Software and represent the 95% confidence interval.
Pre- and post-allotransplantation pathogen detection — the number of allogeneic stem cell transplantations is increasing. This development has intensified the need for rapid and accurate detection, quantification, and identification of pathogens before transplantation to determine whether the donor material is suitable. If the organs, tissue, or stem cells are suitable but there is a pathogen that does not preclude transplantation, for example cytomegalovirus virus (CMV), fast detection and absolute quantification can dictate whether to start preemptive therapy. Furthermore, because allotransplantation recipients are immunosuppressed, there is a need to monitor patients and detect pathogens rapidly.
Pathogens in food — food-borne disease is a major health problem. Rapid and sensitive detection of pathogens in food is required to identify the species and strain, help track down the source of an outbreak, and for routine testing of foods for quality control. Increased sensitivity in detection and measurement of low levels of bacteria such as E. coli and Salmonella will improve surveillance and food safety.
Drug Resistance
The development of drug resistance is a growing problem for the treatment of many infectious diseases. Not only can digital PCR be effective in detecting, profiling, and quantifying drug resistant infections, it can be used to detect heteroresistance, the presence of a small subset of antibiotic-resistant organisms within a large population.
Heteroresistance is common in a number of infectious diseases that are worldwide public health problems. It is thought that heteroresistance can be a precursor to the emergence of antibiotic-resistant strains such as multidrug-resistant Staphylococcus aureus (MRSA) and multiple drug-resistant tuberculosis (MDR-TB).
Current testing methodologies to identify a subset of antibiotic-resistant organisms within a population generally involve culturing bacteria, which can take days. Due to its speed, digital PCR has been shown to be a good tool for identifying heteroresistance, which drug or multidrug resistance is present, and the numbers and proportion of bacteria that are resistant to each drug.
For TB, digital PCR data have been shown to be in agreement with the results from culturing and more accurate than real-time PCR or PCR followed by Sanger sequencing in detecting heteroresistance for isoniazid, rifampin, fluoroquinolone, and aminoglycoside (Pholwot et al. 2013).

VIDEO TESTIMONIAL
Quantifying the Levels of HIV Reservoirs
The Droplet Digital PCR System is used to quantify the level of HIV reservoirs in patients treated with Sangamo BioScience’s unique zinc-finger–based HIV therapy.
Only a small proportion of the total number of microorganisms has been identified, and many remain uncultured or difficult to grow. The advent of PCR resulted in a rapid expansion of the knowledge and study of microorganisms, including those not amenable to culture. The introduction of digital PCR further expands our ability to discover, identify, and analyze microorganisms in small- and large-scale communities.
- Tracking changes in population composition — in microbiomes, changes in environmental conditions may cause shifts in the makeup of a population, both species and their proportions, as well as in the genomes of individual species. To be able to track these changes, analysis must be fast and requires accurate quantification of the composition and structure of a population.
- Reduced sensitivity to PCR inhibitors — one problem for many studies has been that samples taken directly from natural habitats often contain PCR-inhibitory substances such as humic acid. This has been particularly troublesome for real-time PCR, reducing sensitivity and underestimating target gene numbers (Hoshino and Inagaki 2012). Digital PCR helps overcome these problems by eliminating the requirement for reference or standard curves, plus the digital (binary) nature of the data, rather than measurement of the rate of amplification, means that partial inhibition of any DNA polymerase does not affect quantification.
- Cosegregation analysis — digital PCR can be used to determine whether two sequences of DNA are on the same bacterial chromosome by using two fluorescent probes. When droplets are either positive for both probes or negative for both probes after PCR, then the sequences cosegregate. This can be used to characterize a wide array of characteristics such as metabolism, structural components, toxin production, pigmentation, and antibiotic resistance. This linking of sequences can also be used to determine phylogenetic relationships; to link genes to species, digital PCR can be combined with 16s rRNA sequencing.
- Viral-host relationship — digital PCR can be used to link bacterial species to phage markers to study viral-host relationships and lateral phage transfer (Tadmor et al. 2011).
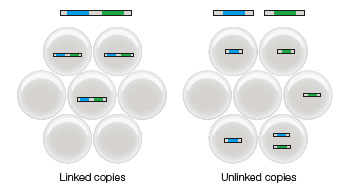
Comparison of expected co-localization for linked and unlinked targets (cosegregation analysis).

VIDEO TESTIMONIAL
Environmental Monitoring for Genetic Targets
Yiping Cao and John Griffith use Bio-Rad's Droplet Digital PCR platform to determine variations in water quality from coastal watersheds to the ocean.
Droplet Digital PCR alone, and integrated with other technologies, will play an increasing role in many areas of microbial research and contribute to improvements in the detection and treatment of infectious diseases as well as provide new methods for basic microbiology studies.
References
Anderson E et al. (2012). Single-copy detection of plasma HIV-1 RNA using droplet digital PCR technology. 19th Annual Conference on Retroviruses and Opportunistic Infections. Paper #679.
Hoshino T and Inagaki F (2012). Molecular quantification of environmental DNA using microfluidics and digital PCR. Syst Appl Microbiol 35, 390–395. PMID: 22840894
Persuad D et al. (2013). Functional HIV cure after very early ART of an infected infant. 20th Annual Conference on Retroviruses and Opportunistic Infections. Paper #48LB.
Pholwot S et al. (2013). Digital PCR to detect and quantify heteroresistance in drug-resistant Mycobacterium tuberculosis. PLoS One 8, e57238. PMID: 23468945
Strain MC et al. (2013). Highly precise measurement of HIV DNA by Droplet Digital PCR. PLoS One 8, e55943. PMID: 23573183
Tadmor AD et al. (2011). Probing individual environmental bacteria for viruses by using microfluidic digital PCR. Science 33, 358–62. PMID: 21719670
Related Content
Droplet Digital PCR Opens New Perspectives in HIV Research (BioRadiations)
