
Overview
Frequently Asked Questions
Learn more about how to use the SEQuoia RNA Library Prep Kit.
The SEQuoia Complete Stranded RNA Library Prep Kit is a high-performance RNA-Seq kit that captures long and short RNAs in a single library, even from limited and low-quality samples. The streamlined workflow minimizes the number of pipetting steps and reduces the overall protocol time to less than 4 hours. The kit allows construction of libraries that are >99% stranded and suitable for next-generation sequencing on Illumina platforms and includes access to SEQSense Analysis Solution and Toolkit, a dedicated secondary data analysis pipeline.
Featuring SEQzyme, a proprietary engineered enzyme that couples cDNA synthesis with adapter addition in a continuous synthesis reaction. The unique enzymatic properties effectively captures all types and sizes of RNA species in a novel enzymatic reaction, significantly improving the diversity and quality of RNA libraries, even from limited or degraded RNA samples.
Features and Benefits
- Captures long and short RNAs in a single prep
- Capable with low sample input requirements
- Validated with FFPE and other low-quality samples
- Full protocol can be completed in less than 4 hours
- Integrated bioinformatics solution
Experience the Power of SEQzyme
Highly Efficient and Streamlined Library Prep Workflow
The unique enzymatic properties of SEQzyme produce robust, high-quality RNA library. The kits are optimized to facilitate efficient library synthesis using a wide range of input RNA (100 pg to 1 μg) and are compatible with automated next-generation sequencing library construction. The streamlined workflow minimizes the number of pipetting steps and greatly reduces the overall protocol time to less than 4 hours.
Diverse Libraries with Low Quality and Limited Input Samples
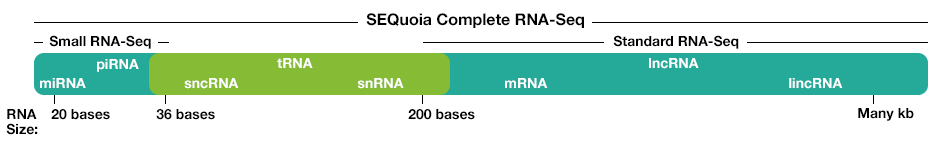
The kit constructs a stranded (>99%) library that captures all RNA biotypes, both long and short RNAs, even from poor quality RNAs (for example, FFPE tissue samples). The feature enables researchers to discover biologically relevant differences in more areas of the transcriptome.
Dedicated Data Analysis Toolkit
SEQSense Analysis Solutions and Toolkit, which is available through a web portal, or can be downloaded and run locally, enables users to trim raw sequencing data and align sequences to a reference genome. The output provides specific quality control metrics to assess the quality of the sequencing data as well as a BAM file, which can be used with tertiary analysis packages to perform advanced analyses.
Applications and Uses
- Differential gene expression and whole transcriptome profiling of high- and low-quality RNA samples
- Sorted or single cell
- Micro-dissection
- Characterization of non-polyadenylated RNAs
- Gene fusion identification
- Single nucleotide variation discovery
- Biomarker discovery
- Splice junctions identification
Included Components
Each kit comes with 2 reagent boxes. Box A includes reagents for RNA fragmentation, cDNA synthesis, and library preparation. The kit is stored at –20°C:
- Reagent A Fragmentation Mix
- Reagent B End Repair Enzyme
- Reagent C Poly(A) Buffer
- Reagent D Poly(A) Polymerase
- Reagent E SEQzyme Mix
- Reagent F Amplification Mix
Box B includes magnetic bead for purification. The kit is stored at 4°C
SEQuoia Dual Indexed Primers are available and can be ordered separately.
SEQuoia Complete RNA Library Preparation Kit FAQs
Kit Contents and Storage
Sample Preparation
RNA Library Amplification
Sequencing and Data Analysis
Configuration Options
|
|
Description |
|
|---|---|---|
| 17005726 | SEQuoia Complete Stranded RNA Library Prep Kit | 24 reactions |
| 17005710 | SEQuoia Complete Stranded RNA Library Prep Kit | 96 reactions |
| 12011928 | SEQuoia Dual Indexed Primers Set, 12 vials of Unique Dual Indexes | 96 reactions |
| 12011930 | SEQuoia Dual Indexed Primers Plate, one 96-well plate of Unique Dual Indexes | 96 reactions |
Specifications
Ordering

A high-performance RNA sequencing (RNA-Seq) kit that captures long and short RNAs in a single library, even from limited and low-quality samples. The kit provides all of the components required to make an RNA-Seq library for Illumina platforms. The size of the kit is for 24 reactions.

A high-performance RNA sequencing (RNA-Seq) kit that captures long and short RNAs in a single library, even from limited and low-quality samples. The kit provides all of the components required to make an RNA-Seq library for Illumina platforms. The size of the kit is for 96 reactions

For use with SEQuoia Complete Stranded RNA Library Prep Kits. Contains 12 unique dual indexed primers supplied in 12 separate vials, for 96 reactions

Accessories

Pkg of 20, bag of 12 x 8 thin-wall polypropylene tube strips and 12 x 8-cap strips for PCR, natural (1,920 tubes and 1,920 caps)

Pkg of 20, bag of 8 x 12 thin-wall polypropylene tube strips and 8 x 12-cap strips for PCR, natural (1,920 tubes and 1,920 caps)




