On This Page |
What Is CRISPR? | Hands-On Lab Activities | Paper Activities | CRISPR Videos and Other Resources |
On This Page |
What Is CRISPR? | Hands-On Lab Activities | Paper Activities | CRISPR Videos and Other Resources |
What Is CRISPR?
-

A Microbial Immune System and a Gene Editing Technology
CRISPR gene editing technology is the result of decades of research into genome sequences and the adaptive immune system of bacteria immune system of bacteria and archaea. Read below to learn what CRISPR sequences are and how they work with Cas enzymes to incorporate snippets of viral DNA into the bacterial genome. Then extend your understanding to how these elements work with host cell DNA repair mechanisms in the CRISPR-Cas9 gene editing workflow.
CRISPR-Cas — a Microbial Immune System
CRISPR gene editing technology is the result of decades of research into genome sequences and the adaptive immune system of bacteria and archaea. The following are some highlights:

A CRISPR region within a microbial genome

This video from The Franklin Institute summarizes how research in multiple areas came together to give the world CRISPR-Cas9 technology.
- In the late 1980s and early 1990s, researchers including Francisco Mojica noticed unusual repetitive DNA sequences in prokaryotic genomes. These sequences were short, palindromic (read the same forward and backward), and repeated many times with "spacer" sequences between the repeats. These unique sequences were called "clustered regularly interspaced short palindromic repeats" (CRISPR) sequences
- The function of CRISPR sequences remained a mystery until the early 2000s, when Alexander Bolotin and colleagues identified the CRISPR-associated (Cas) protein called Cas9 and proposed its role as a DNA-cutting enzyme in the bacterial immune system
- In 2007, Philippe Horvath and his team demonstrated that CRISPR sequences work with Cas proteins to provide adaptive immunity against viruses
CRISPR sequences and Cas enzymes work together to incorporate snippets of viral DNA into the bacterial genome. This allows bacteria to recognize and defend against future infections in three phases:
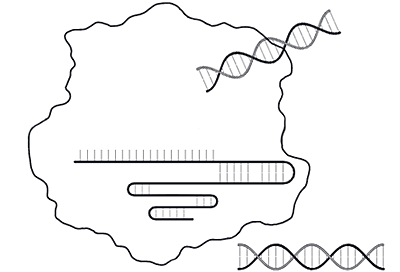
- Cutting and capture — When bacteria are infected by a virus, they use their CRISPR system to cut up the invading viral DNA and insert pieces of it (spacers) into their own genome as a "memory" of the infection
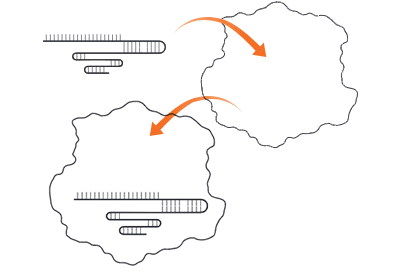
- Monitoring — Bacteria transcribe the spacer DNA into CRISPR RNA (crRNA); Cas9 and the crRNA form a complex and monitor the cell for any DNA sequence complementary to the RNA
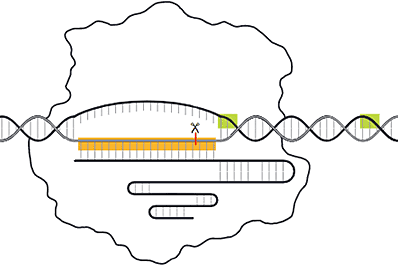
- Defense — If matching (viral) DNA is encountered, the crRNA and RNA-Cas9 complex binds to and cuts the viral DNA to prevent it from replicating. This halts the viral infection
Using this system, bacteria can collect sequences from many different infecting viruses to create a "library." Since the CRISPR sequence is contained in genomic DNA, it is passed on to each generation, and the library continues to change and adapt to more common threats over time.
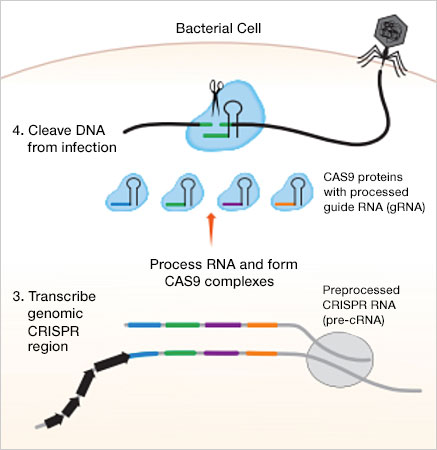
The CRISPR-Cas9 microbial defense system. 1. The Cas1-Cas2 enzymes of the microbe recognize and cut out a segment of foreign DNA. 2. The Cas1-Cas2 enzymes insert the DNA segment into the CRISPR region of the bacterial genome as a spacer. 3. A spacer sequence is transcribed and then linked to a Cas9 protein. 4. Upon reinfection by the same invader, the CRISPR-Cas9 complex can recognize the foreign DNA sequence and cut it to prevent complete reinfection.
CRISPR-Cas9 — A Gene Editing System
In 2012, Jennifer Doudna and Emmanuelle Charpentier demonstrated that the CRISPR-Cas9 system could be repurposed to edit DNA in living organisms. They earned the 2020 Nobel Prize for Chemistry for their work. In 2017, Feng Zhang and colleagues demonstrated the use of CRISPR-Cas9 to edit mammalian genomes.
The CRISPR-Cas9 system uses modified components of the bacterial CRISPR system to direct target-specific cutting of double-stranded DNA. DNA repair mechanisms then take over to fix the break in a manner that modifies the genetic sequence that has been cut.
Step 1: Cutting the DNA

Visit our YouTube CRISPR Gene Editing playlist for videos that depict the CRISPR-Cas9 method for genome editing.
- Cas9 enzyme (Cas9) — an endonuclease that cuts both strands of DNA at a specific site. Multiple types of Cas enzymes are found in nature, but Cas9 is commonly used in the laboratory
- Single guide RNA (sgRNA) — an engineered RNA that forms a complex with Cas9. The sgRNA is a fusion of two regions that occur as separate RNAs in nature:
- Guiding region — part of the CRISPR RNA or crRNA in nature, a 20-nucleotide region that is complementary to the target region and defines the target DNA sequence that Cas9 cuts. Scientists customize this sequence for their own targets
- Scaffold region — the trans-activating CRISPR RNA or tracrRNA in nature, this region forms a multi-hairpin loop structure (scaffold) that binds in a crevice of the Cas9 protein
- Protospacer adjacent motif (PAM) — required for Cas9 function, this sequence motif is immediately downstream of the target sequence. Cas9 recognizes the PAM sequence 5’-NGG, where N can be any nucleotide (A, T, C, or G). When Cas9 binds the PAM, it separates the DNA strands of the adjacent sequence to allow binding of the sgRNA. If the sgRNA is complementary to that sequence, Cas9 cuts the DNA
Step 2: Repairing the Break to Engineer the Change
Researchers can use the cell's own DNA repair machinery to modify, insert, or delete a nucleotide sequence. The repair can happen in two ways:

This video from Science Communication Lab explains homology directed repair (HDR).
- Non-homologous end joining (NHEJ) — enzymes reconnect the ends of the double-stranded break back together. This process may randomly insert or delete one or more bases and can cause mutations that can disrupt gene function or expression
- Homology directed repair (HDR) — proteins patch the break using donor template DNA. Researchers design the donor template DNA that may include a desired sequence flanked on both sides by "homology arms" that match the sequence upstream and downstream of the cut. A complementary DNA strand is created during the repair
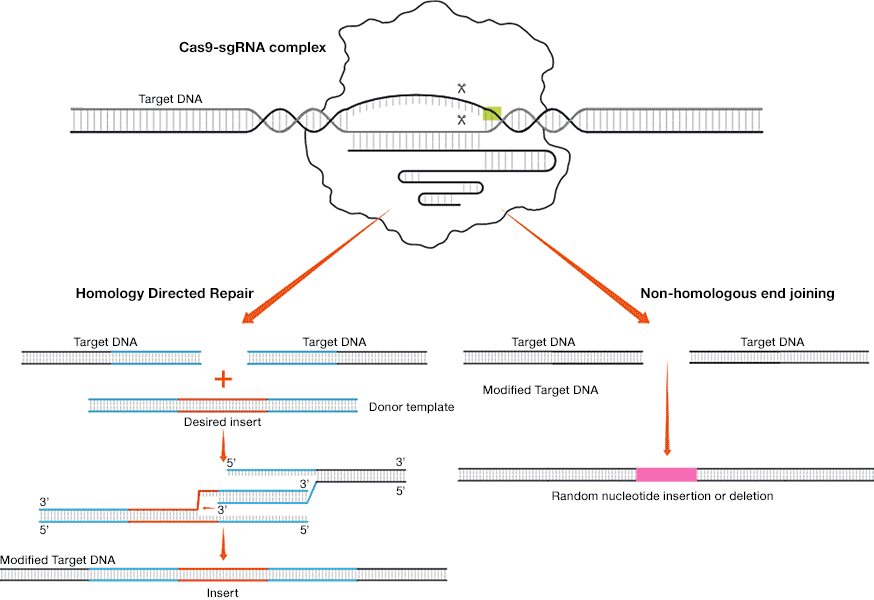
DNA repair via homology directed repair and non-homologous end joining (Scroll right to view image)
Hands-On CRISPR Gene Editing Lab for the Classroom

Out of the Blue CRISPR and Genotyping Extension Kits
Use CRISPR-Cas9 gene editing to edit a bacterial chromosome! These lab kits use familiar — and safe — reagents, techniques, and organisms to bring students to the cutting edge of molecular biology. Using carefully designed bacterial strains and plasmids, students see and learn how the CRISPR-Cas9 system works with homology-directed repair (HDR) to introduce a stop codon into the chromosomal lacZ gene of E. coli.
Order Kits
CRISPR Paper Model & Bioinformatics Activity
-
Image

CRISPR Paper Model
Use this free CRISPR paper model to enhance understanding of how the CRISPR-Cas9 system directs precise cuts of DNA.
-
Image

CRISPR Bioinformatics Activity
Use this activity to explore the design of Cas9 target sites in the human genome and determine risk for off-target effects.
CRISPR Videos & Other Resources

YouTube CRISPR Gene Editing Playlist
Access technique videos, recorded webinars, and perspectives about CRISPR from leading experts.

PowerPoint Presentation for Classroom Use
Use this student-facing slide deck to help guide your students through the CRISPR-Cas9 gene editing lab activity.

Out of the Blue CRISPR Kit Student Activity Video Quick Guide
Use this overview video to prepare for the CRISPR gene editing lab for the Out of the Blue CRISPR Kit.

CRISPR in the Classroom
Looking to add CRISPR into your curriculum? We've got you covered with the essentials you'll need to introduce this cutting-edge topic to your students.

Webinars
New to CRISPR? Join us at a webinar or view on-demand CRISPR-related webinars.

Workshops
Join us at a hands-on workshop or attend an upcoming webinar. Gain insights on how to teach about CRISPR and its many applications.

CRISPR Poster (Free)
The poster features medical breakthroughs enabled by CRISPR gene editing technology, as well as a timeline of this and other Nobel prizes earned by women in science.
Add to Cart
CRISPR Infographic (Free)
This infographic provides an overview of the CRISPR timeline and the regulatory, legal and ethical debates it has led to – great topics of discussion in your classroom!
Add to Cart