
The ProteOn™ XPR36 protein interaction array system enables label-free quantitative analysis of biomolecular interactions in real time using surface plasmon resonance (SPR) technology. The interactions are monitored over time by flowing an analyte in a microfluidic channel over a ligand immobilized on a sensor chip and detecting the binding of the analyte to the ligand by measuring changes in the propagation of electromagnetic waves at the sensor surface. In this section, you will learn more about SPR technology, the heart of the ProteOn XPR36 system, and the types of biological research that can benefit from using the ProteOn system.
In the following, we describe how SPR technology works and what kinds of biomolecular interactions can be studied using the ProteOn system, including applications such as the determination of binding affinity and specificity, binding kinetics, binding site stoichiometry, and binding mechanisms.
Related Topics: ProteOn Stystem/Detection and Sensor Chips.
Page Contents
Surface plasmon resonance is an optical phenomenon that is used in the ProteOn system to monitor the binding of any two unlabeled molecules in real time. The SPR signal is based on changes in the refractive index at the surface of a gold sensor chip as an analyte flows in a microfluidic channel and binds to a ligand immobilized on the sensor chip. Monitoring the change in the SPR signal over time produces a sensorgram, a plot of the binding response versus time. Fitting the sensorgram data to a suitable kinetic binding model allows for the calculation of kinetic parameters such as the association (ka) and dissociation (kd) rate constants.
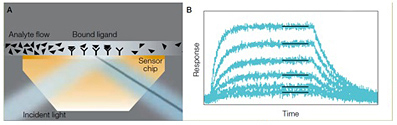
Schematic illustration of the conversion of shifts in SPR angle into sensorgrams. A, as analyte binds to ligand molecules on the sensor chip surface, the intensity minimum (shadow) produced by the surface plasmon resonance effect undergoes an angular shift. This intensity shift is measured in real time for 36 interaction spots and 42 interspot references; B, each of the resulting sensorgrams is fitted to an appropriate mathematical model to quantitatively characterize the interaction (here shown for one set of six).
In addition to obtaining kinetic data, the ProteOn system can be used to investigate the specificity and thermodynamics of binding, screen compound libraries for affinity, or characterize epitopes of an antigen. The novel XPR™ technology of the ProteOn system is a unique application of SPR that uses enhanced microfluidics and a 6 × 6 interaction array on the ProteOn sensor chip to characterize the binding of multiple analytes to multiple ligands, in a single step, on a single chip.
ProteOn sensor chips come coated with a modified alginate polymer that provides a biomimetic surface for ligand immobilization. The hydrophilic nature of the alginate coating prevents surface denaturation of immobilized ligands, blocks non-specific analyte binding, and allows analyte-ligand binding to occur much as it does in solution. The alginate polymer can be modified with several different functional groups to facilitate different immobilization chemistries. The basic ProteOn sensor chip comes functionalized with carboxylic acid groups that react with the ligand via amide coupling of surface-exposed amine groups. More specialized ProteOn sensor chips such as the NLC and HTG/HTE chips enable site-specific immobilization of biotinylated or polyhistidine-tagged ligands, respectively, and can be used in situations where standard amine coupling sensor chips are not appropriate. In addition, experiment-specific capturing agents such as secondary antibodies can be amine-coupled to the general-binding sensor chip to ensure the selective capture of specific ligands in a uniform orientation. The wide variety of sensor chips available for the ProteOn system provides users with the flexibility to immobilize or capture diverse targets of interest. Making SPR technology easy to use through offline solutions (with the NLC, HTG, and HTE chips) or customization through online solutions (with the amine coupling chips), the ProteOn chips provide high-quality, reproducible SPR assays for a broad range of applications.

ProteOn sensor chips.
Videos
Learn how the ProteOn can be applied to many important applications of the drug discovery workflow.
Learn how the ProteOn system's design provides data you can trust through unique referencing options.
Learn in more detail how the ProteOn system uses SPR technology to address important experimental questions.
TEST
General ProteOn Literature
| 6411 | ProteOn XPR36 Quantikinetics: Antibody Concentration and Detailed Kinetic Analysis in a Single Experimental Cycle | Click to download |
| 5390 | ProteOn — PIA XPR36 Brochure | Click to download |
| 5538 | ProteOn XPR36 — Analyzing Protein Interaction Array System Featured Article Reprint | Click to download |
| 5413 | ProteOn XPR36 Hardware — 36 Interactions on a Single Chip: Label-Free, in Real-Time PIS | Click to download |
| 5627 | ProteOn Manager Software PIS | Click to download |
| 5404 | ProteOn Sensor Chips — Application-Specific Surface Chemistries = Optimized Ligand Activty PIS | Click to download |
| 5410 | ProteOn Protocol Development Kits PIS | Click to download |
| 5409 | Protein Interaction Analysis — ProteOn XPR36 System Ordering Information Sheet | Click to download |
| 6449 | Novel Liposome-Capture Surface Chemistries to Analyze Drug-Lipid Interaction using the ProteOn XPR36 System, Rev A | Click to download |
| 6414 | ProteOn XPR36 Experimental Design and Application Guide, Rev B | Click to download |
Large Molecule Protein-Interaction Analysis
| 3172 | ProteOn PIA — Rapid and Efficient Determination of Kinetic Rate Constants using the ProteOn XPR36 Protein Interaction Array System Tech Note | Click to download |
| 5412 | ProteOn App Guide — Antibody Characterization and Development Using the ProteOn XPR36 PIA System App Guide PIS | Click to download |
| 5540 | ProteOn — PIA Screening, Ranking, and Epitope Mapping of Anti-Human IL-9 Supernatants TN | Click to download |
| 5368 | ProteOn PIA — Analysis of Multiple Protein-Protein Interactions Using the ProteOn XPR Protein Interaction Array System Tech Note | Click to download |
| 5449 | Protein Interaction Analysis Applications of the ProteOn NLC Sensor Chip: Antibody-Antigen, DNA-Protein-Protein Interactions Tech Note | Click to download |
| 5358 | ProteOn PIA — Mechanisms of Protein Binding: Double-Mutant Cycle Analysis Using the ProteOn XPR36 System Tech Note | Click to download |
| 5820 | ProteOn — Rapid Screening and Selection of Optimal Antibody Capturing Agents Using the ProteOn XPR36 Protein Interaction Array System | Click to download |
| 5360 | ProteOn PIA — Rapid and Detailed Analysis of Muliple Antigen-Antibody Pairs Using the ProteOn XPR36 Protein Interaction Array System Tech Note | Click to download |
| 5367 | ProteOn PIA — Rapid Optimization of Immobilization and Binding Conditions for Kinetic Analysis of Protein-Protein Interactions using the ProteOn XPR36 PI Array System Tech Note | Click to download |
Small Molecule Analysis
| 5965 | Rapid High-Throughput Screening of Protein Kinase Inhibitors Using the ProteOn PIA System | Click to download |
| 5797 | Protein Interaction — Rapid Assay Development and Optimization for Small Molecule Drug Discovery Tech Note | Click to download |
| 5679 | ProteOn — Applications of the ProteOn GLH Sensor Chip: Interaction Between Proteins and Small Molecules | Click to download |
| 5960 | High-Throughput Profiling of Kinase Inhibitors Selectivity Using the ProteOn XPR36 Protein Interaction Array System | Click to download |
| 5846 | Determining the Binding Kinetics of HIV-1 Nucleocapsid Protein to Six Densities of Oligonucleotide Using the ProteOn XPR36 Protein Interaction Array System TN | Click to download |
| 5822 | How to Perform Excluded Volume Correction on the ProteOn XPR36 Protein Interaction System PG | Click to download |

