
Bio-Rad's ProteOn™ XPR36 protein interaction array system uses surface plasmon resonance (SPR) technology to detect and monitor biomolecular interactions in real time for label-free interaction analysis. Binding events are detected by monitoring the change in the SPR signal, which is proportional to changes in mass at the sensor chip surface over time as an analyte flows through a microfluidic channel and interacts with a target immobilized to the sensor chip. The ProteOn XPR36 system can be used to monitor many different biomolecular interactions including those between antibodies and antigens, enzymes and their substrates/inhibitors, small molecules and drug discovery targets, and whole cells and lipid membranes. SPR technology is flexible; the applications for SPR are vast, and experimental design can be tailored to individual needs. An additional advantage of the ProteOn XPR36 instrument is that unlike some traditional SPR biosensors, the unique XPR™ technology and 6 × 6 interaction array allow for the simultaneous measurement of up to 36 biomolecular interactions. XPR technology greatly speeds time to results for traditional kinetic measurements by enabling the patented One-shot Kinetics™ approach, whereby up to six targets are immobilized to the sensor chip surface followed by a single, orthogonal injection of six unique analytes. In addition to traditional kinetic measurements, SPR technology can be used to qualitatively assess biomolecular interaction properties, monitor the quality and/or concentration of biologics, and investigate binding thermodynamics.
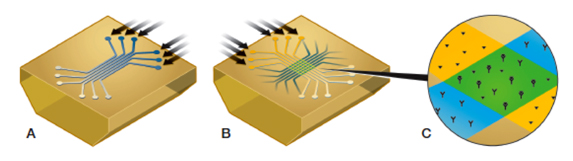
Generation of the 6 × 6 ligand-analyte interaction array. A, six ligands are immobilized in six parallel ligand channels (blue); B, six analyte samples are injected into six analyte channels perpendicular to the six ligand channels (yellow); C, detail of a single ligand-analyte interaction spot (green) showing the positions of the two interspot references (yellow).
Learn more about label-free SPR technology and how the ProteOn XPR36 system can be used for diverse applications in label-free interaction analysis including kinetic screening, validation, protein quantitation, and assay development and experimental optimization.
Related Topics: Large and Small Molecule Screening by SPR, Label-Free Interaction Validation and Characterization, and SPR Assay Development and Experimental Optimization.
Page Contents
The versatility of the microfluidics in the ProteOn system make it an ideal complement to the drug discovery and development workflow for target screening and characterization studies. The ProteOn system is capable of monitoring up to 36 unique biomolecular interactions simultaneously in a variety of configurations depending on the experimental design:
1 × 1 — full optimization is used to quickly determine optimal assay conditions.
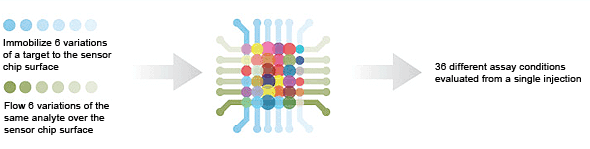
6 × 1 — kinetic screening is used to measure kinetics under different conditions in the same run (One-shot Kinetics).
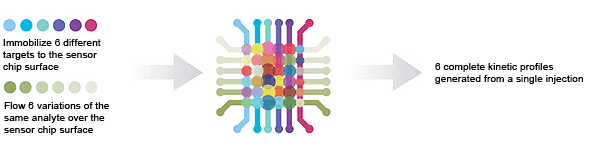
6 x 6 — single-point screening is used to screen libraries for interactions of interest.

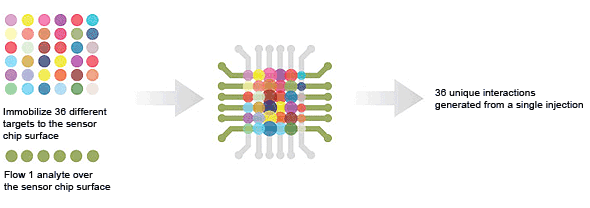
36 × 1 — target screening is used to screen an analyte against 36 targets in a single injection.
The ProteOn XPR36 system can also be used in downstream processing such as protein quantitation for quality control, as SPR can be used to monitor the concentration of small molecules and biologics in crude or impure samples.
Large and Small Molecule Screening
The versatility of the ProteOn XPR36 platform expands the capabilities of SPR analysis, making this technology ideal for use in several stages of the drug development pipeline. With the rise of antibody therapeutics, there is a need for fast and accurate determination of the affinity of candidate antibodies to their targets. SPR can be used for both quantitative large and small molecule screening and qualitative relative ranking of antibody therapeutics (binding or no binding) during drug development. Small molecules can also be screened for activity and desirable adsorption, distribution, metabolism, and excretion (ADME) properties using the ProteOn system. This ability enables the identification of undesirable compounds earlier in the drug discovery process, before considerable time and effort are invested in costly clinical trials.
Validation and Characterization
Unlike endpoint assays such as isothermal calorimetry that measure a binding response after reaching equilibrium, SPR allows for the detection of binding events in real time and the accurate measurement of both association (ka) and dissociation (kd) rate constants. This additional information can be helpful in designing potent agonists/antagonists with a fast ka and slow kd or for understanding quantitative structure-activity relationships of small molecules. The unique One-shot Kinetics approach of the ProteOn XPR36 system allows for robust kinetic characterization from a single analyte injection (Yousef 2007). In addition, biomolecular interaction thermodynamics can be characterized by using the ProteOn system to quantify kinetics at different temperatures.
Assay Development and Experimental Optimization
Due to the vast array of applications for SPR, label-free interaction protocols for the ProteOn XPR36 must be tailored to specific experimental needs to obtain high-quality SPR data. The With the deposition of six or more ligands on one sensor chip, the ProteOn XPR36 system’s unique 6 × 6 interaction array allows for flexibility in the design of biomolecular interaction assays, streamlining assay development. Interrogating these different ligands with six separate analytes facilitates experimental optimization by enabling the real-timed detection of up to 36 different biomolecular interaction events simultaneously.
Videos
Learn how the ProteOn can be applied to many important applications of the drug discovery workflow.
Learn how the ProteOn system's design provides data you can trust through unique referencing options.
Learn in more detail how the ProteOn system uses SPR technology to address important experimental questions.
TEST
General ProteOn Literature
| 5390 | ProteOn — PIA XPR36 Brochure | Click to download |
| 5538 | ProteOn XPR36 — Analyzing Protein Interaction Array System Featured Article Reprint | Click to download |
| 5413 | ProteOn XPR36 Hardware — 36 Interactions on a Single Chip: Label-Free, in Real-Time Product Information Sheet | Click to download |
| 5627 | ProteOn Manager Software Product Information Sheet | Click to download |
| 5404 | ProteOn Sensor Chips — Application-Specific Surface Chemistries = Optimized Ligand Activty Product Information Sheet | Click to download |
| 5410 | ProteOn Protocol Development Kits Product Information Sheet | Click to download |
| 5409 | Protein Interaction Analysis — ProteOn XPR36 System Ordering Information Sheet | Click to download |
| 6449 | Novel Liposome-Capture Surface Chemistries to Analyze Drug-Lipid Interaction using the ProteOn XPR36 System, Rev A | Click to download |
| 6414 | ProteOn XPR36 Experimental Design and Application Guide, Rev B | Click to download |
Large Molecule Protein-Interaction Analysis
| 3172 | ProteOn PIA — Rapid and Efficient Determination of Kinetic Rate Constants using the ProteOn XPR36 Protein Interaction Array System | Click to download |
| 5412 | ProteOn App Guide — Antibody Characterization and Development Using the ProteOn XPR36 PIA System Application Guide Product Information Sheet | Click to download |
| 5540 | ProteOn — PIA Screening, Ranking, and Epitope Mapping of Anti-Human IL-9 Supernatants | Click to download |
| 5368 | ProteOn PIA — Analysis of Multiple Protein-Protein Interactions Using the ProteOn XPR Protein Interaction Array System | Click to download |
| 5449 | Protein Interaction Analysis Applications of the ProteOn NLC Sensor Chip: Antibody-Antigen, DNA-Protein-Protein Interactions | Click to download |
| 5358 | ProteOn PIA — Mechanisms of Protein Binding: Double-Mutant Cycle Analysis Using the ProteOn XPR36 System Tech Note | Click to download |
| 5820 | ProteOn — Rapid Screening and Selection of Optimal Antibody Capturing Agents Using the ProteOn XPR36 Protein Interaction Array System | Click to download |
| 5360 | ProteOn PIA — Rapid and Detailed Analysis of Muliple Antigen-Antibody Pairs Using the ProteOn XPR36 Protein Interaction Array System | Click to download |
| 5367 | ProteOn PIA — Rapid Optimization of Immobilization and Binding Conditions for Kinetic Analysis of Protein-Protein Interactions using the ProteOn XPR36 PI Array System | Click to download |
Small Molecule Analysis
| 5965 | Rapid High-Throughput Screening of Protein Kinase Inhibitors Using the ProteOn PIA System | Click to download |
| 5797 | Protein Interaction — Rapid Assay Development and Optimization for Small Molecule Drug Discovery | Click to download |
| 5679 | ProteOn — Applications of the ProteOn GLH Sensor Chip: Interaction Between Proteins and Small Molecules | Click to download |
| 5960 | High-Throughput Profiling of Kinase Inhibitors Selectivity Using the ProteOn XPR36 Protein Interaction Array System | Click to download |
| 5846 | Determining the Binding Kinetics of HIV-1 Nucleocapsid Protein to Six Densities of Oligonucleotide Using the ProteOn XPR36 Protein Interaction Array System | Click to download |
| 5822 | How to Perform Excluded Volume Correction on the ProteOn XPR36 Protein Interaction System Product Guide | Click to download |

