Gene expression analysis is most simply described as the study of the way genes are transcribed to synthesize functional gene products — functional RNA species or protein products. The study of gene regulation provides insights into normal cellular processes, such as differentiation, and abnormal or pathological processes.
Related Topics: Gene Cloning and Analysis, Mutational Analysis, Epigenetics and Chromatin Structure, and Nucleic Acid Analysis
Page Contents
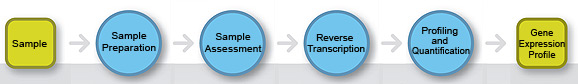
Gene expression workflow.
The following text-based list provides full accessible navigation for all steps shown in the gene expression workflow image.
Researchers may perform gene expression analysis at any one of several different levels at which gene expression is regulated: transcriptional, post-transcriptional, translational, and post-translational
protein modification.
Transcription, the process of creating a complementary RNA copy of a DNA sequence, can be regulated in a variety of ways. Transcriptional regulation processes are the most commonly studied and manipulated in typical gene expression analysis experiments.
The binding of regulatory proteins to DNA binding sites is the most direct method by which transcription is naturally modulated. Alternatively, regulatory processes can also interact with the transcriptional machinery of a cell. More recently, the influence of epigenetic regulation, such as the effect of variable DNA methylation on gene expression, has been uncovered as a powerful tool for gene expression profiling. Varying degrees of methylation are known to affect chromatin folding and strongly affect accessibility of genes to active transcription.
Following transcription, eukaryotic RNA is typically spliced to remove noncoding intron sequences and capped with a poly(A) tail. At this post-transcriptional level, RNA stability has a significant effect on functional gene expression, that is, the production of functional protein. Small interfering RNA (siRNA) consists of double-stranded nucleic acid molecules that are participants in the RNA interference pathway, in which the expression of specific genes is modulated (typically by decreasing activity). Precisely how this modulation is accomplished is not yet fully understood. A growing field of gene expression analysis is in the area of microRNAs (miRNAs), short RNA molecules that also act as eukaryotic post-transcriptional regulators and gene silencing agents.
Researchers studying gene expression employ a wide variety of molecular biology techniques and experimental methods. Gene expression analysis studies can be broadly divided into four areas: RNA expression, promoter analysis, protein expression, and post-translational modification.
RNA Expression
- Northern blotting — steady-state levels of mRNA are directly quantitated by electrophoresis and transfer to a membrane followed by incubation with specific probes. The RNA-probe complexes can be detected using a variety of different chemistries or radionuclide labeling. This relatively laborious technique was the first tool used to measure RNA levels
- DNA microarrays — an array of oligonucleotide probes bound to a chip surface enables gene expression profiling of many genes in response to a condition. Labeled cDNA from a sample is hybridized to complementary probe sequences on the chip, and strongly associated complexes are identified optically. Gene expression profiling is often a first step in a gene expression analysis workflow, investigating changes in the expression profile of a whole system or examining the effects of mutations in biological systems
- Real-Time PCR — steady-state levels of mRNA are quantitated by reverse transcription of the RNA to cDNA followed by quantitative PCR (qPCR) on the cDNA. The amount of each specific target is determined by measuring the increase in fluorescence signal from DNA-binding dyes or probes during successive rounds of enzyme-mediated amplification. This precise, versatile tool is used to investigate mutations (including insertions, deletions, and single-nucleotide polymorphisms (SNPs)), identify DNA modifications (such as methylation), confirm results from northern blotting or microarrays, and conduct gene expression profiling. Expression levels can be measured relative to other genes (relative quantification) or against a standard (absolute quantification). Real-time PCR is the gold standard in nucleic acid quantification because of its accuracy and sensitivity. Real-time PCR can be used to quantitate mRNA or miRNA expression following conversion to cDNA or to quantitate genomic DNA directly to investigate transcriptional activity
Promoter Analysis
- Expression of reporter genes/promoter fusions in host cells — promoter activity (transcription rate) is measured in vivo by introducing fusions of various promoter sequences with a gene encoding a product that can be readily measured to monitor activity levels
- In vitro transcription (nuclear run-on assays) — transcription rates are measured by incubating isolated cell nuclei with labeled nucleotides, hybridizing the resultant product to a membrane (slot blot), and then exposing this to film or other imaging media
- Gel shift assays — also called electrophoretic mobility shift assays, these are used to study protein-DNA or protein-RNA interactions. DNA or RNA fragments that are tightly associated with proteins (such as transcription factors) migrate more slowly in an agarose or polyacrylamide gel (showing a positional shift). Identifying the associated sequences provides insight into gene regulation
- Chromatin immunoprecipitation (ChIP) — protein-binding regions of DNA can be identified in vivo. In living cells, DNA and protein are chemically cross-linked, and the resulting complex is precipitated by antibody-coated beads (immunoprecipitation). Following protein digestion and DNA purification, the sequences of the precipitated DNA are determined
Protein Expression
- Western blotting — quantification of relative expression levels for specific proteins is accomplished by electrophoretically separating extracted cell proteins, transferring them to a membrane, and then probing the bound proteins with antibodies (targeted to antigens of interest) that are subsequently detected using various chemistries or radiolabelling
- 2-D Gel Electrophoresis — protein expression profiling is achieved by separating a complex mixture of proteins in two dimensions and then staining to detect differences at the whole-proteome level
- Immunoassays — proteins are quantitated in solution using antibodies that are bound to color-coded beads (as in the Bio-Plex supension array system) or immobilized to a surface (ELISA), which is subsequently probed with an antibody suspension and is typically detected using a chromogenic or fluorogenic reporter
Posttranslational Modification Analysis
- Immunoassays — levels of protein phosphorylation and other post-translational modifications are detected using antibodies that are specific for these adducts
- Mass spectrometry — proteins and their modifications are identified based on their mass
Bustin SA et al. (2009). The MIQE guidelines: Minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55, 611–622. PMID: 19246619
Lefever S et al. (2009). RDML: Structured language and reporting guidelines for real-time quantitative PCR data. Nucleic Acids Res 37, 2065–2069. PMID: 19223324
Vandesompele J et al. (2002). Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3, research0034.1. PMID: 12184808
Guénin S et al. (2009). Normalization of qRT-PCR data: The necessity of adopting a systematic, experimental conditions-specific, validation of references. J Exp Bot 60, 487–493. PMID: 19264760
Kitchen RR et al. (2010). Statistical aspects of quantitative real-time PCR experiment design. Methods 50, 231–236. PMID: 20109551
Yuan JS et al. (2006). Statistical analysis of real-time PCR data. BMC Bioinformatics 7, 85. PMID: 16504059
Related Content
Videos
This tutorial demonstrates how to perform a simple experiment to quickly identify the best electroporation conditions for your primary cell transfection project. The video also shows how optimal conditions identified using 96-well electroporation plates can be used with standard electroporation cuvettes.
This tutorial highlights the main components and features of the Gene Pulser Xcell system. It provides information about system installation and the setup of electroporation experiments, including important troubleshooting tips and answers to frequently asked questions. Ordering information for system components and accessories is also provided.


