On This Page | Importance of Data Analysis | Data Acquisition and Analysis | Gene Expression Analysis | BR.io Cloud Connectivity | Related Products | Resources |
Real-time quantitative polymerase-chain-reaction (qPCR) is a standard technique in most research laboratories performing gene expression. qPCR data analysis is a crucial part of a gene expression experiment, and has led to the development of several key methods. The sections which follow provide an overview of the key quantification strategies used in qPCR for gene expression.
The Importance of Data Analysis in Gene Expression
Gene expression analysis by real-time qPCR has been a key enabler of a routine and robust approach for measuring gene expression in genes of interest, as well as monitoring biomarkers. This section will provide the key features of qPCR data analysis and describe examples of common methods to analyze data from a qPCR assay.
How to Evaluate Amplification Curves from a qPCR Assay
Structure of an amplification curve can be defined in terms of the phases described below.

Figure 1. Amplification curve. The different phases of the qPCR assay reaction. Baseline occurs from cycles 0 through 10, where the initial concentration of template is low. At this level, the fluorescence intensity is too low to be detected and only background signal occurs. Exponential: once target yield attains the detection threshold, the reaction is followed through its exponential phase. Linear: template concentration increases, leading to a reduction in available DNA polymerase concentration, and the reaction rate decreases. Plateau: reaction is at maximal yield.
Introduction to Data Acquisition and Analysis: Basic Calculations using Cq
The quantitative analysis of RT-qPCR is obtained through analysis of the quantification of cycle (Cq) values. The Cq value has been given many different names including:
- Ct — threshold cycle
- Cp — crossing point
- TOP — take-off point
According to MiQE guidelines, Cq is the correct terminology.
How to Identify the Cq Value for qPCR
The Quantification Cycle, Cq, is the cycle number at which the fluorescence first rises above the threshold level. In the diagram below, note where Cq is defined in relation to the baseline level and the beginning of the exponential phase of the reaction curve.
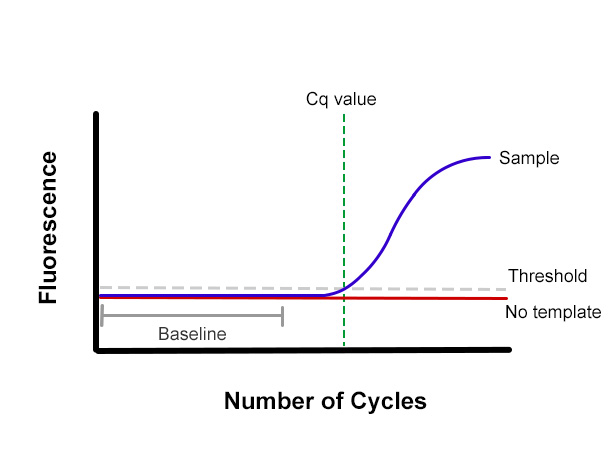
How to Analyze Data to Determine Cq
A set of procedures and data normalization is required before performing quantification analysis:
Baseline Correction
To avoid variation from background signal caused by external factors unrelated to the samples (e.g. plastic ware, light leaking, fluorescence probe not quenched, etc…), it is recommended to select the fluorescence intensity from the first cycles, usually from cycles 5-15 and identify a constant and linear component of background fluorescence. This information is used to define the baseline for analysis.
Threshold Setting
The quantification zone should be selected which exceeds the detection limit of the PCR thermocycler being used. Usually the number of cycles needed is directly proportional to the sample copy number. Instead of using the intensity of minimum fluorescence detected by the instrument, a quantification zone with higher fluorescence intensity is selected and a threshold defined for this area.
Using the following guidelines can help define a correct threshold:
- Lies in the amplification log phase, and avoids the plateau phase
- Lies significantly over the background baseline
- Lies within a log phase range where sample amplification plots are parallel
To see a real-world example of analysis for gene expression, review the example listed below.
Gene Expression Analysis
The most common application of real-time PCR is the study of gene expression. Measuring gene expression is of fundamental importance in man areas of contemporary biomedical research, ranging from basic science to practical applications in industry and medicine. In most gene expression studies, researchers are interested in determining the relative gene expression levels in test vs. control samples. For example, a researcher might be interested in the expression of a particular gene in cancerous vs. normal tissue. In this section, general guidelines and a sample gene expression experiment are presented to demonstrate how to perform gene expression analysis using real-time PCR.
Experimental Design
A sample gene expression analysis using a multiplex Taqman assay is presented in the following sections. In this example, we're interested in the relative expression of three genes in the polyamine biosynthesis pathway, ornithine decarboxylase (ODC), ODC antizyme (OAZ), and antizyme inhibitor (AZI), in two different samples, sample A and sample B. Due to possible pipetting errors and initial quantification errors of the input RNA, the amount of starting cDNA in the different real-time reactions may be different. In this example, the expression of a reference gene, β-actin, was chosen as the normalizer to control for any difference in cDNA input amount. The following steps were performed to determine the relative expression level of ODC, OAZ, and AZI in the two different samples:
- RNA was isolated from sample A and sample B.
- RNA was reverse transcribed into cDNA.
- The amount of the target genes (ODC, OAZ, and AZI) and the reference gene (β-actin) was determined in each of the cDNA samples using a multiplex qPCR assay.
- Data were analyzed and the relative expression of each of the target genes in the two samples was calculated.
Experimentation
To determine the expression of the three target genes, ODC, OAZ, and AZI, and the reference gene, β-actin, in samples A and B, these four genes were assessed in a multiplex two-step RT-qPCR assay.
Reaction Components for Multiplex Assay
The multiplex assay contained the following components in a 50 µl reaction
- 1/10 of a reverse transcription reaction that used 100 ng of total RNA
- 300 nM each primer
- 200 nM each probe
- 200 µM each dNTP
- 5 mM MgCl2
- 3.75 U iTaq DNA polymerase
Cycling Protocol
The cycling program used is shown below:
| Step | Description | Temperature °C | Time | Number of Cycles |
|---|---|---|---|---|
| 1 | Polymerase Activation and DNA Denaturation | 95 | 2 min | 1 |
| 2 | Amplification: Denaturation | 95 | 10 sec | 35–45 |
| 3 | Amplification: Annealing, Extension and Plate Read | 55 | 45 sec |
Learn more about specific features in Bio-Rad's CFX Maestro Software for qPCR data analysis »
.
CFX Opus Cloud Connectivity: BR.io
Get started using
BR.io today!
The CFX Opus System seamlessly integrates with Bio-Rad's new cloud platform, BR.io, enabling you to get the most out of your instrument and minimize time at the bench. BR.io can be accessed from any internet connection using a Safari or Chrome web browser, and there is no software installation required.
BR.io cloud connectivity eliminates the need for a dedicated computer connected to the instrument and provides new capabilities.
Remotely set up CFX Opus runs, and start directly from the instrument
Monitor instruments and run progress remotely
Automatically transfer and store data in the cloud
Access and analyze data from anywhere
BR.io Cloud Platform Tutorial Video Series
Learn how to use the BR.io cloud platform with a CFX Opus Real-Time PCR System. Videos in this series:
- 1. Introduction to the BR.io Cloud Platform
- 2. Getting Started in the BR.io Cloud Platform
- 3. Linking a CFX Opus Real-Time PCR System to Your BR.io Cloud Platform Account
- 4. Setting Up a CFX Opus Protocol in BR.io
- 5. Setting Up and Performing a CFX Opus Real-Time PCR System Run Using the BR.io Cloud Platform
- 6. Reviewing and Exporting Data from a CFX Opus Real-Time PCR System Run Using the BR.io Cloud Platform

Questions about the CFX Opus Cloud qPCR System?
Related Products
qPCR Detection Systems
These systems deliver sensitive, reliable detection of both singleplex and multiplex real-time PCR reactions.
qPCR Analysis Software
CFX Maestro Software is designed to streamline data collection, visualization, and analysis. Precision Melt Analysis software enables high resolution melt for SNP Genotyping and detection of small indels.
Resources
Amplification and Reagents and Plastics Brochure
An overview of PCR reagents and plastics with info on specifications, performance data, and more.
PCR Amplification Guide
A guide for RT-PCR on key concepts, experimental designs, analyses, and product recommendations.









