Green Fluorescent Protein (GFP) is a protein from the jellyfish Aequorea victoria. It emits a bright green fluorescence when exposed to ultraviolet light. In research labs, scientists attach the gene that encodes GFP to other genes to create a “tag” that glows. This helps them track gene expression and label specific proteins, cells, or tissues.
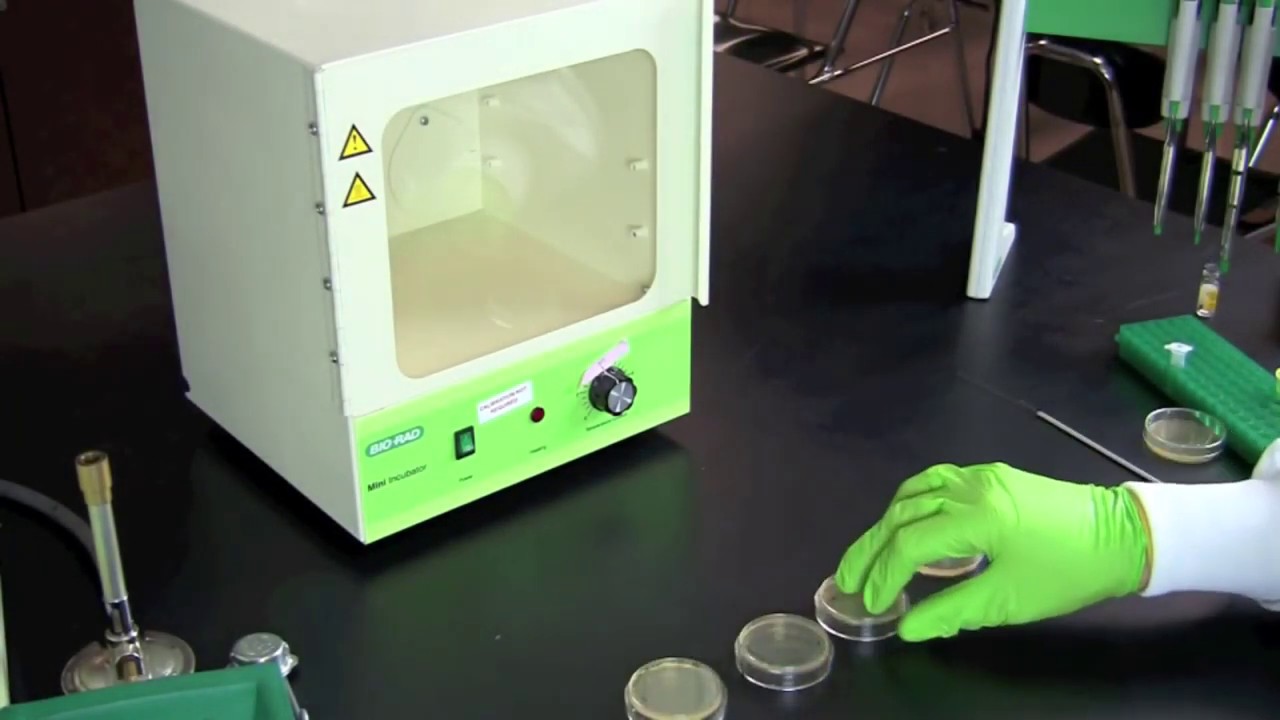
In science classrooms, teachers can use GFP technology to offer students an unforgettable hands-on lab activity to explore the central dogma, gene expression and regulation, bacterial transformation, protein separation, and the biomanufacturing process. Bio-Rad pGLO Bacterial Transformation Kits use the pGLO plasmid, which contains the gene encoding GFP. The kits enable students to perform bacterial transformation and express GFP in a non-virulent laboratory strain of Escherichia coli (E. coli).
This page provides background information and a selection of free presentations, videos, case studies, and other resources to help teachers bring these concepts to light.
pGLO Plasmid
The pGLO plasmid is designed to enable (1) replication of the plasmid in bacteria, (2) selection of transformants using ampicillin resistance, and (3) regulation of GFP expression. Bacteria transformed with the pGLO plasmid are selected by ampicillin resistance and then induced to express GFP to glow fluorescent green under UV light!
The essential sequences in the pGLO plasmid include the following:
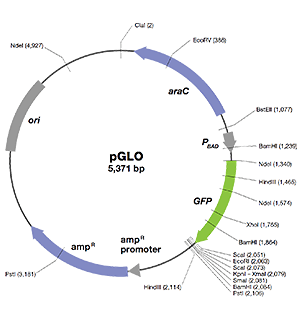
- GFP — jellyfish gene that encodes green fluorescent protein (GFP)
- ori — origin of pGLO plasmid DNA replication (essential for making more copies of the plasmid)
- bla — gene that encodes β-lactamase, an enzyme that breaks down the antibiotic ampicillin; transformants expressing the bla gene can be selected by adding ampicillin in the growth medium
- pBAD promoter — binds AraC-arabinose and promotes RNA polymerase binding and transcription of the GFP gene
- araC — gene that encodes a regulatory protein that binds to the pBAD promoter; only when arabinose binds to the AraC protein is the production of GFP switched on
- Multiple cloning site — a region containing restriction sites (NdeI, HindIII, EcoRI, etc.), sequences that permit the insertion or deletion of a gene of interest
-
Image

pGLO Map & Sequence
Download a PDF containing the pGLO plasmid map and its complete sequence.
-
Image

pGLO Sequence
Download a text (.txt) file of the complete pGLO plasmid sequence.
Bacterial Transformation
Scientists use bacterial transformation to introduce foreign DNA into bacterial cells, which enables them to manipulate and study genes, gene function, and gene expression. The pGLO Bacterial Transformation Kits use the CaCl2/heat shock method to enable bacterial transformation with the pGLO plasmid.
A standard technique used in research laboratories and in biomanufacturing, the CaCl2/heat shock protocol can accommodate virtually any classroom laboratory. It can be completed with no additional laboratory equipment — insulated cups, warm water, and ice are all that is required. Alternatively, students can use this lab as an opportunity to gain experience with general lab equipment such as micropipets, water bath or heat blocks, and laboratory incubators.
The elements of the heat shock protocol and how they work with the pGLO plasmid are explained in the PowerPoint files that accompany each kit.
Gene Regulation
Gene expression is carefully regulated to allow organisms to adapt to environmental conditions and prevent wasteful production of proteins. Regulation often occurs at the level of transcription from DNA into RNA, specifically at the promoter, where RNA polymerase binds the DNA and begins transcription of the gene. The pGLO plasmid is a fantastic tool for teaching gene regulation in a hands-on and engaging way.
-

- pGLO LB
-

- pGLO LB/amp
-

+ pGLO LB/amp
-

+ pGLO LB/amp/ara
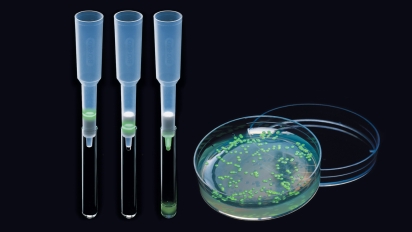
Results of a pGLO bacterial transformation experiment. Controls that were incubated with no plasmid (-pGLO) grow as a lawn in the absence of ampicillin (LB plate) and do not grow at all in the presence of ampicillin (LB/amp). Transformants grown on ampicillin (LB/amp) grow as colonies but do not show GFP fluorescence; those grown in the presence of both amp and ara (LB/amp/ara) do glow green under UV light.
In bacteria, groups of related genes are often clustered together and transcribed into RNA from one promoter. These clusters of genes controlled by a single promoter are called operons. The bacterial genes encoding the enzymes needed to metabolize the simple sugar arabinose are a perfect example: Three genes that encode the digestive enzymes involved in breaking down arabinose (araB, araA, and araD) are clustered together in arabinose operon 3, and all depend on initiation of transcription from a single promoter, pBAD. Transcription requires the simultaneous presence of RNA polymerase, a DNA-binding protein called AraC, and arabinose.



With the pGLO plasmid, then, the GFP gene is transcribed only when arabinose is present. The AraC protein will bind to the arabinose, allowing GFP transcription to mRNA to take place. This mRNA is then translated into green fluorescent protein (GFP), a process known as gene expression. As the cells produce GFP, they fluoresce a bright green.
This is an excellent example of the central dogma of molecular biology in action: DNA > RNA > protein > trait.
In the absence of arabinose, the GFP gene is not expressed, and the bacterial colonies exhibit a wild-type (natural) phenotype, appearing white with no fluorescence. By plating pGLO transformants on media that contain or don’t contain arabinose, students can see a very clear example of gene regulation in action.
The pGLO Bacterial Transformation Kits are carefully designed to include controls that facilitate teaching and learning about the regulation of gene expression. For more details, refer to the PowerPoint files that accompany each kit (posted also in this page).
Green Fluorescent Protein (GFP)

View this informative video about GFP from Phospho Biomedical Animation and other videos available in the YouTube pGLO Bacterial Transformation Playlist.
In nature, GFP fluoresces in the deep-sea jellyfish, Aequorea victoria. GFP has a barrel structure surrounding a central alpha helix that contains the fluorophore. It can be used as an example for discussions of protein secondary structure, parallel and anti-parallel beta sheets, and the use of genes and proteins in biotechnology.
GFP does not require enzymes to fluoresce. Its small size (27 kDa) allows it to be fused to other proteins without disrupting their function but to enable cellular localization and tracking. Osamu Shimomura, Martin Chalfie, and Roger Tsien earned a Nobel Prize in 2008 for their development of GFP as a tool that continues to be used in pioneering new discoveries.
GFP extracted from transformed bacteria retains its fluorescent properties even when separated by chromatography.
Remarkably, GFP retains its fluorescent properties when expressed in
The pGLO Bacterial Transformation and GFP Kit family includes the SDS-PAGE Extension and GFP Chromatography Kits. The unique fluorescence of GFP allows real-time monitoring of extraction and purification, key processes used in biotechnology to produce and purify designer proteins with commercial or research value.
pGLO Bacterial Transformation & GFP Kits

Bio-Rad’s pGLO Bacterial Transformation and GFP Kits offer engaging and unforgettable student lab activities in which students engineer bacteria to express the Green Fluorescent Protein (GFP) and glow green under UV light. These kits enable hands-on learning about the central dogma, gene expression and regulation, genetic engineering, protein separation, and the biomanufacturing process.
Learn More
PowerPoint Presentations
-
Image

pGLO Bacterial Transformation (PPT 4.4 MB)
Use this student-facing presentation to facilitate use of the classic pGLO Bacterial Transformation Kit.
-
Image

pGLO Bacterial Transformation for General Biology (PPT 22.1 MB)
Use this student-facing presentation to facilitate use of the pGLO Bacterial Transformation Kit for General Biology.
-
Image

pGLO Bacterial Transformation for AP Biology (PPT 41.4 MB)
Use this student-facing presentation to facilitate use of the pGLO Bacterial Transformation Kit for AP Biology.
-
Image

Bring Inquiry Into Your Classroom with the pGLO Plasmid (PPT 9.06 MB)
Learn how to use pGLO bacterial transformation to illustrate the science and engineering practices described in the NGSS framework.
-
Image

GFP Purification — Electrophoresis and Chromatography (PPT 9.63 MB)
Learn how to use SDS-PAGE and/or chromatography to purify glowing GFP from your pGLO transformants.
pGLO Videos & Webinars
-
Image

YouTube pGLO Bacterial Transformation Playlist
A collection of videos and recorded webinars to enhance understanding of bacterial transformation, GFP, gene regulation, and genetic engineering. Includes useful lab preparation videos, too!
-
pGLO Bacterial Transformation Student Activity Video Quick Guide
Prepare students for the lab portion of the follow along with the pGLO Bacterial Transformation Kit and pGLO Transformation and Inquiry Kit for AP Biology.
Case Studies
-
Image

Case Study: A Role for Bacterial Transformation in Controlling Malaria Transmission (PDF 3.3 MB)
A student-facing extension that is also useful for AP Biology exam prep, this case study connects bacterial transformation with the global fight against malaria.
-
Image

Case Study: Hacking the Gut Microbiome (PDF 1.5 MB)
A student-facing extension that is also useful for AP Biology exam prep, this case study connects bacterial transformation with the role of the microbiome in health and disease.
Other pGLO Resources and Ideas
-
Image

The Biotech Universe Poster
Add color to your wall with this free, full-length poster that highlights the pGLO plasmid and flow of biological information from DNA to RNA to protein to trait.
Add to Cart -
Image

pGLO Art
Have your students make artwork using pGLO transformants! These fun pGLO plates were made by students in Valerie May’s AP Biology course at The Woodstock Academy in Woodstock, Connecticut.
-
Image

Microbes in Space
Read about how high school students designed experiments for the International Space Station with
Bio-Rad pGLO Bacterial Transformation Kits.