
Conventional electrophoresis uses a single electrical field to cause biomolecules to migrate through a matrix according to its mass-to-charge ratio; the migration distance of the biomolecule is indicative of its mass or size (Klotz and Zimm 1972). Conventional electrophoresis can effectively separate DNA fragments up to ~20 kb. However, larger fragments will comigrate and appear as a large band at the top of the gel when imaged. In 1984, Schwartz and Cantor invented pulsed field gel electrophoresis (PFGE) to overcome this problem. PFGE resolves DNA by alternating the electrical field between spatially distinct pairs of electrodes. This technique results in the separation of DNA fragments of up to ~10 Mb by their reorientation and movement at different speeds through the pores of an agarose gel. This section provides an overview of pulsed field gel electrophoresis and describes factors to be considered during sample preparation for PFGE.
Related Topics: Horizontal Electrophoresis, and CHEF Solutions.
Page Contents
PFGE stemmed from the observation that DNA molecules elongate upon application of an electric field and return to an unelongated state upon removal of the electric field; this relaxation rate is dependent on the size of the DNA.
When the orientation of the electric field is changed during electrophoresis, the DNA molecules must return to their elongated form prior to reorientation, thus affecting the migration rate. This effect can be used to greatly extend the size range over which electrophoretic DNA separations are possible.
When the electrical field is applied to the gel, the DNA molecules elongate in the direction of the electrical field. The first electrical field is then switched to the second field according to the run specifications. The DNA must change conformation and reorient before it can migrate in the direction of this field.
As long as the alternating fields are equal with respect to the voltage and pulse duration, the DNA will migrate in a straight path down the gel (see below).
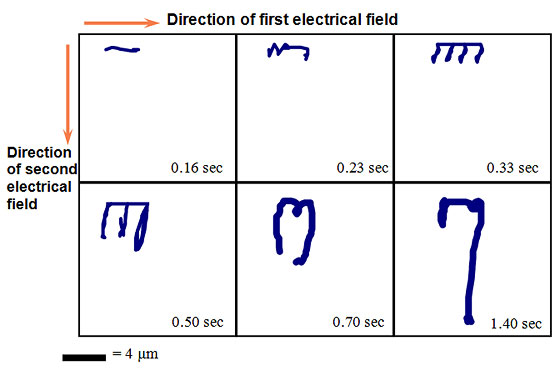
Time-lapse representation of DNA molecules undergoing PFGE.
The large size of DNA molecules to be separated by PFGE imposes certain constraints on sample preparation and handling. High molecular weight DNA is easily cleaved through shearing and imparts very high solution viscosity. For these reasons, DNA samples for PFGE are generally prepared by embedding in gel medium. Cellular source material is suspended in low gelling agarose and the gelled suspension is poured into molds. All subsequent manipulations (for example, cell lysis, protein removal, and restriction digestion) are performed by diffusing reagents into the resultant gel plugs. The processed gel plugs are then carefully loaded into wells of an agarose gel used for PFGE.
Find more information on troubleshooting PFGE gels, plug preparation, preparing and running a gel, image analysis, PFGE size standards, restriction enzyme digestion, and critical parameters in optimizing sample resolution in the Protocols section below.
- CHEF (clamped homogeneous electrical field) technology resolves DNA over a wide range of molecular weights in a straight lane; it employs the principles of contour-clamped electrophoresis to generate homogenous electrical fields. More information on CHEF
- PACE (programmable autonomously controlled electrodes) technology allows users to select the angle of electrophoretic pulsing optimal for the desired size range
- DR (dynamic regulation) is the electronics design by which each of the 24 electrodes is regulated; CHEF-DR® systems are capable of compensating for changes in buffer conductivity or gel size, preventing these changes from affecting the reproducibility of results
- FIGE (field inversion gel electrophoresis) is used for rapid sample resolution in the 100 bp to 250 kb size range; in FIGE the electrical field is fixed at 1 angle (180°) and is inverted in the forward and reverse directions
- AFIGE (asymmetric field inversion gel electrophoresis) is a further refinement of the FIGE technology; AFIGE applies a different voltage to the forward direction electrical field than to the reverse direction electrical field, which optimizes the sample resolution in the FIGE size range
Videos
Documents
TEST
| 6223 | General Considerations for Troubleshooting PFGE Gels | Click to download |
| 6224 | Plug Preparation Protocol | Click to download |
| 6225 | Restriction Enzyme Digestion Protocol | Click to download |
| 6226 | Preparing and Running a Gel | Click to download |
| 6227 | Image Analysis | Click to download |
| 6228 | Pulsed Field Gel Electrophoresis Standards | Click to download |
| 6229 | Optimizing Sample Resolution | Click to download |

