Protein solubilization is the process of breaking interactions involved in protein aggregation, which include disulfide bonds, hydrogen bonds, van der Waals forces, ionic interactions, and hydrophobic interactions. If these interactions are not prevented, proteins can aggregate or precipitate, resulting in artifacts or sample loss. For successful electrophoretic separation, proteins must be well solubilized. This section describes various agents used for protein solubilization, their method of action, and their compatibility to different types of samples. It also provides troubleshooting tips.
Cell lysis and protein solubilization are ideally carried out in the sample buffer that is recommended for the particular electrophoresis technique, especially when native electrophoresis is the method of choice. If this is not possible, proteins must be prepared in sample solubilization solutions that typically contain a number of compounds, including chaotropic agents, detergents, reducing agents, buffers, salts, and ampholytes. These are chosen from a list of compounds that meet the requirements, both electrically and chemically, for compatibility with the electrophoretic technique being used. In these cases, the sample will have to be diluted with concentrated electrophoresis sample buffer to yield a 1x final buffer concentration. Sample solubilization methods for 2-D electrophoresis applications are discussed in the 2-D Electrophoresis section.
Related Sections: Cell Disruption, Removal of Interfering Substances, and Sample Quantitation.
Page Contents
Detergents are classified as nonionic, zwitterionic, anionic, and cationic, and they disrupt hydrophobic interactions between and within proteins. Some proteins, especially membrane proteins, require detergents for solubilization during isolation as well as to maintain solubility. Nonionic detergents such as NP-40 and Triton X-100 are not very effective in solubilizing hydrophobic proteins; zwitterionic detergents such as CHAPS and sulfobetaines (for example, SB 3–10 or ASB 14) provide higher solubilization efficiency, especially for integral membrane proteins (Luche et al, 2003). Sample preparation for SDS-PAGE commonly uses the anionic detergent sodium dodecyl sulfate (SDS), which is unparalleled in its ability to efficiently and rapidly solubilize proteins.
Thiol reducing agents such as 2-mercaptoethanol βME and dithiothreitol (DTT) disrupt intramolecular and intermolecular disulfide bonds and are used to achieve complete protein unfolding and to maintain proteins in their fully reduced states (see figure below).
βME is volatile, evaporates from solutions, and reduces protein disulfide bonds by disulfide exchange. There is equilibrium between free thiols and disulfides, so βME is used in large excess in sample buffers to drive the equilibrium reaction toward completion. If the concentration of βME drops and proteins reoxidize, they may yield fuzzy or spurious artifactual bands upon gel electrophoresis of the sample.
DTT is less volatile and is altered during the disulfide reduction reaction to form a ring structure form its original straight chain. The equilibrium favors protein reduction, so lower concentrations of DTT are needed (higher concentrations are recommended for proteins with large numbers of disulfide bonds).
Phosphines such as tributylphosphine (TBP) and triscarboxyethylphosphine (TCEP)* offer an alternative to thiols as reducing agents because they can be used at lower concentration and over a wider pH range than the sulfhydryl reductants.
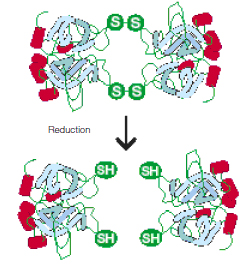
Reduction of proteins with DTT.
*TCEP is included in Bio-Rad’s XT sample buffers. Though TCEP can be added to SDS-PAGE sample buffer, it must first be neutralized with NaOH; otherwise it will hydrolyze proteins.
Chaotropic compounds such as urea disrupt hydrogen bonds and hydrophobic interactions both between and within proteins. When used at high concentrations, they destroy secondary protein structure and bring otherwise-insoluable proteins into solution.
Urea and substituted urea such as thiourea improve solubilization of hydrophobic proteins.
Samples containing urea and thiourea can be used in SDS-PAGE when diluted with SDS-PAGE sample buffer. In this case, the protein solution may not be heated above 37°C, since urea and thiourea can hydrolyze to cyanate and thiocyanate, respectively, and modify amino groups on proteins (carbamylation), giving rise to artifactual charge heterogeneity.
Both pH and ionic strength influence protein solubility; therefore, buffer choice is important, especially when native electrophoresis conditions are required. Many proteins are more soluble at higher pH, so Tris base is often included to elevate the pH. However, proteins differ in their solubility at different pH values, so the use of different buffers can extract different sets of proteins. The choice of buffer and pH of the sample preparation solution can strongly influence which proteins are successfully solubilized.
Even in the presence of detergents, some proteins have stringent salt requirements to maintain their solubility, but salt should be present only if it is an absolute requirement. Excess salt in SDS-PAGE samples causes fuzzy bands and narrowing of gel lanes toward the bottom of the gel. If the ionic strength is very high, no visible bands will appear in the lower part of the gel (a vertical streak will appear instead) and the dye front will be wavy instead of straight.
| Problem | Cause | Solution |
| Laemmli sample buffer turns yellow | Sample buffer is too acidic | Add Tris base until buffer turns blue again |
| Sample very viscous | High DNA or carbohydrate content |
Fragment DNA with ultrasonic waves during cell lysis and protein solubilization Add endonucleases like Benzonase Precipitate protein with TCA/acetone (ReadyPrep 2-D cleanup kit) to diminish carbohydrate content |
Documents
TEST
| 6194 | Protein Sample Generation General Tips | Click to download |
| 6195 | Protein Sample Preparation (Human Tissue) | Click to download |
| 6196 | Protein Sample Preparation (Mammalian Tissue) | Click to download |
| 6197 | Protein Sample Preparation (Plant Leaves) | Click to download |
| 6198 | Protein Sample Preparation (Microbial Cultures) | Click to download |
| 6199 | Buffer Formulations | Click to download |
