
Protein extraction and sample cleanup are the most important steps to ensure optimal resolution and reduce variability of your 2-D gels. 2-D PAGE success depends on sample purity.
Interfering substances that can negatively impact SDS-PAGE and 2-DGE include salts, detergents, denaturants, or organic solvents, so it is crucial to eliminate these contaminants prior to analysis. Highly viscous samples indicate high DNA and/or carbohydrate content, which may interfere with separations. In addition, solutions at extreme pH values (for example, fractions from ion exchange chromatography) diminish the separation power of most electrophoresis techniques.
This section provides tips for total protein extraction, removal of contaminants such as salt, lipids, detergents, phenolic compounds, nucleic acids, polysaccharides, and plant-specific compounds, and for disulfide bond reduction.
Related Topics: Cell Disruption, Protein Solubilization, and Protein Fractionation and Depletion.
Page Contents
Bio-Rad's protein sample preparation products may be used with virtually all sample types. These products are based on well-understood purification and fractionation principles, and may be applied individually or in combination for effective sample preparation.
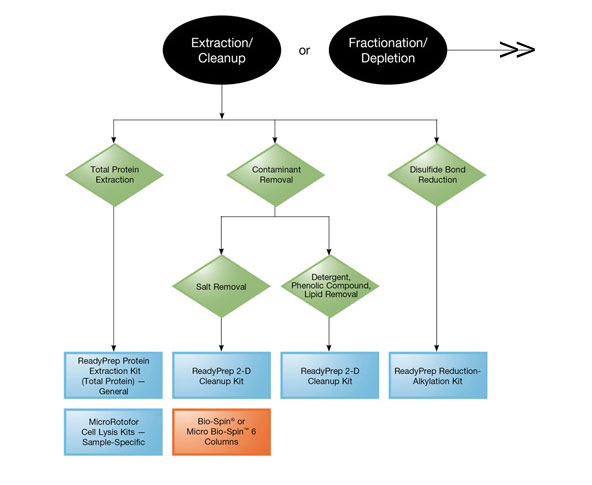
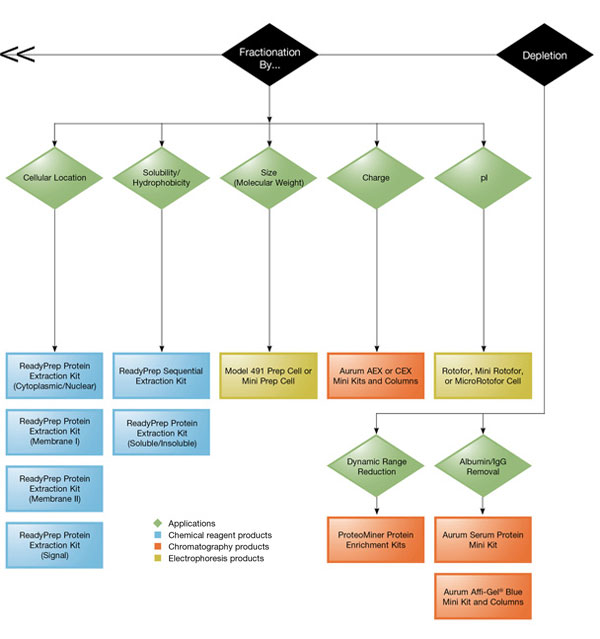
Product Selection Guide (extraction and cleanup).
Bio-Rad offers ReadyPrep™ protein extraction kits and the MicroRotofor™ cell lysis kits, which provide cell lysis and protein extraction protocols that are tailored to the specific needs of different sample sources.
ReadyPrep Protein Extraction Kit (Total Protein) — includes a strongly chaotropic extraction solution containing the zwitterionic detergent ASB-14. It generates protein samples that can be applied directly to IEF and 2-D gel electrophoresis.
MicroRotofor Cell Lysis Kits — four kits tailored to different organisms are available. All four kits are based on the same chaotropic protein solubilization buffer (PSB), which contains nondetergent sulfobetaine 201 (NDSB 201) along with urea, thiourea, and CHAPS for effective solubilization of most proteins. The kits generate total protein samples that are ready to be applied to SDS-PAGE, IEF, and 2-D gel electrophoresis. Different sample types have different requirements for effective cell disruption, and all four kits combine PSB with other elements to accommodate these specific needs.
- MicroRotofor cell lysis kit (mammal) — uses maceration and solubilization (up to a 100 mg sample) using ReadyPrep mini grinders and PSB. The kit is intended for use with samples from mammalian sources, but it may also be applied to other animal tissues
- MicroRotofor cell lysis kit (bacteria) — for use with both gram-negative and gram-positive bacterial cultures. It employs enzymatic digestion using lysozyme followed by solubilization into PSB
- MicroRotofor cell lysis kit (plant) — for use with soft plant tissues and cultured cells. It employs maceration (up to a 1 g sample) and solubilization into PSB. A ReadyPrep™ 2-D cleanup kit is provided to enable removal of plant-derived phenolics, which may interfere with IEF
- MicroRotofor cell lysis kit (yeast) — for use with yeast cultures (~60 µl wet cell pellet). It employs enzymatic digestion of the cell wall with lyticase followed by solubilization into PSB
Commercially available kits designed for removal of salts, high-abundance proteins, and other contaminants incorporate procedures such as affinity and size exclusion chromatography to improve resolution of 2-D gels.
Salt
Removal of salts reduces streaking and improves reproducibility of 2-D gels (see figure below). Use either buffer exchange (desalting) or protein precipitation (which can also help concentrate the sample if needed).
Methods to remove salts from sample:
- Dilution — only works if the protein concentration is high in the starting sample
- Dialysis — some proteins may adsorb to the dialysis membrane, however a detergent usually prevents this; often used under native condition where denaturation must be avoided
- Protein precipitation — the most versatile method to selectively separate proteins from contaminants consists of protein precipitation by trichloroacetic acid (TCA)/acetone followed by resolubilization in electrophoresis sample buffer
- Buffer Exchange — size exclusion chromatography is another effective method for removing salts, detergents, and other contaminants
| Before | After |
 |
 |
Salt removal using the ReadyPrep 2-D cleanup kit. E. coli extracts containing 1 M NaCl were electrophoresed before and after treatment with the 2-D cleanup kit. The samples were focused using 11 cm ReadyStrip™ pH 3–10 IPG strips, then run on Criterion™ 8–16% Tris-HCl precast gels for the second dimension.
Bio-Rad offers specialized products such as ReadyPrep™ 2-D cleanup kit for the removal of ionic contaminants such as detergents, lipids, and phenolic compounds from protein samples, and Bio-Spin® and Micro Bio-Spin™ 6 Columns for the removal of salt and contaminants.

Bio-Spin column.
Lipids, Detergents, and Phenolic Compounds
Ionic contaminants such as lipids, detergents, and phenolic compounds interfere with isoelectric focusing of proteins resulting in poor 2-D gel resolution and reproducibility.
Methods to remove lipids, detergents, and phenolic compounds from sample:
- Detergent — in general, the detergent present in the sample buffer, such as CHAPS, will be sufficient enough to solubilize lipids and prevent interference
- Solvent extraction/precipitation — acetone precipitation and in rare cases chloroform/methanol extraction will remove lipids and enable subsequent electrophoresis; One problem with the use of organic solvents is that the proteins are often precipitated with the solvents, and may be hard or impossible to resolubilize. However, some samples may be best solubilized with an organic solvent
| Before | After |
 |
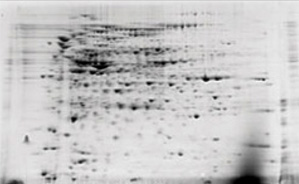 |
Detergent removal using the ReadyPrep 2-D cleanup kit. E. coli extracts containing 1% SDS were electrophoresed before and after treatment with the ReadyPrep 2-D cleanup kit. The samples were focused using 11 cm ReadyStrip pH 3–10 IPG strips, then run on Criterion 8–16% Tris-HCl precast gels for the second dimension.
Nucleic Acids
The presence of nucleic acids, especially DNA, interferes with separation of proteins by IEF. Under denaturing conditions, DNA complexes are dissociated and markedly increase the viscosity of the solution, which inhibits protein entry and slows migration in the immobilized gradient gel (IPG). In addition, DNA binds to proteins in the sample and causes artifactual migration and streaking.
Methods to remove nucleic acids from sample:
- Enzymatic digestion — simplest method for removal of DNA; sample preparation can be achieved in a single step, by the addition of the enzyme prior to loading the first-dimension IPG. Adding endonuclease to the sample after solubilization at high pH (40 mM Tris) allows efficient digestion of nucleic acids while minimizing the action of contaminating proteases
- Ultra-centrifugation — large nucleic acids will sediment and polycations such as polyethyleneimine are added to disrupt protein-nucleic acid interactions; protein loss is high
- Mechanical disruption — ultrasonic probes and bead mills are effective tools for shearing nucleic acids into fragments that are too small to interfere with IEF
Polysaccharides
The presence of polysaccharides interferes with separation of proteins by IEF. The effects are similar to nucleic acids, but are far more difficult to remove from samples.
Methods to remove polysaccharides from sample:
- Ultra-centrifugation — large polysaccharides will sediment and complexing agents, such as spermine or polyethlyeneimine, are added to disrupt polysaccharide interactions; there is a high risk of protein loss
- Solvent extraction/precipitation — acetone precipitation and other solvent extractions will remove polysaccharides and enable subsequent electrophoresis (i.e., TCA, TCA/acetone, ammonium sulfate, phenol/ammonium acetate, and organic solvent extraction)
Plant-Specific Compounds
Plant-specific compounds such as tannins, lignins, and chlorophyll can cause severe disruption of electrophoresis, in particular IEF.
Methods to remove plant-specific compounds from sample:
- Organic solvent extraction/precipitation — acetone and/or TCA/acetone precipitation efficiently removes these interfering compounds present in plants and enable subsequent electrophoresis
Disulfide bond formation is problematic for basic proteins because of their increased rate of formation in an alkaline environment. Protein solubility is also limited, which can interfere with resolution. Moreover, many reducing agents become negatively charged during IEF and migrate off the IPG strip, thus allowing disulfide bonds to re-form. Reducing disulfide bonds ensures proper protein migration and often yields better resolution of protein spots, fewer streaks, and greater reproducibility in 2-D separations.
Reduction-Alkylation
Reducing agents are commonly used during sample preparation to cleave disulfide bond crosslinks within proteins and between subunits. Reduction in disulfide bonds decreases streaking and improves reproducibility of 2-D gels (see figure below).
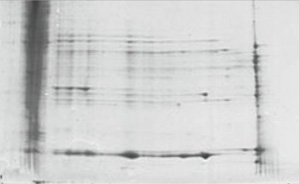
| Reduced using DTT | Reduced Using ReadyPrep Kit |
 |
 |
Disulfide bond removal using the ReadyPrep reduction-alkylation kit. Protein samples were reduced using either 50 mM DTT in rehydration/sample buffer or the reduction-alkylation kit. Both samples were applied by cup loading onto 11 cm ReadyStrip pH 7–10 IPG strips and focused for the first dimension, then run on Criterion 8–16% Tris-HCl precast gels for the second dimension.
Bio-Rad offers the following reducing and alkylating reagents:
- Dithiothreitol (DTT) — a commonly used general sulfhydryl reducing agent. DTT has a pI of about 8 and may not remain effective throughout IEF. It may become negatively charged during IEF and migrate off the IPG strip, thus allowing disulfide bonds to re-form
- Tributylphosphine (TBP) — a phosphine reducing agent that can be used at lower concentrations than DTT and is active over a wider pH range. TBP remains with proteins throughout focusing
- Iodoacetamide — an alkylating agent that can be used following treatment with a reducing agent to permanently modify protein sulfhydryls, preventing proteins from aggregating, and precipitating due to oxidative crosslinking. This ensures that proteins remain soluble throughout electrophoresis and produces a spot pattern with more spots, fewer streaks, and greater reproducibility, particularly in the basic regions of 2-D gels
TBP and iodoacetamide are provided as components of the ReadyPrep Reduction-Alkylation kit , which also provides a complete protocol for performing irreversible reduction and alkylation reactions.
Videos
Horizontal streaks in 2-D gels are the result of poor protein separation during isoelectric focusing (IEF). This tutorial discusses the most common sources of horizontal streaks in 2-D gels: problems with sample preparation and inappropriate IEF conditions.
Documents
TEST
| Number | Description | Options |
|---|---|---|
| 6221 | Sample Quantitation (RC DC Protein Assay) | Click to download |
| 6220 | Solubilization | Click to download |

