Droplet Digital™ PCR (ddPCR™) is a breakthrough technology that provides ultrasensitive nucleic acid detection and absolute quantification. It is highly effective for resolving low abundance targets, such as allelic or structural variants, that are below the level of detection of other platforms. This technology is now approved for use in in vitro diagnostics.
With advanced multiplexing offerings, ddPCR technology maximizes the number of targets analyzed per well without sacrificing performance.
Featured PRODUCTQX600 Droplet Digital PCR System
Since its introduction, ddPCR technology has advanced critical fields of research through unmatched sensitivity, precision, and absolute quantitation. Now the QX600 System provides advanced, 6-color multiplexing to maximize the number of biomarkers interrogated per well, allowing the preservation of precious samples and reducing lab burden.
Digital PCR Assays & Kits

Bio-Rad offers a comprehensive portfolio of Digital PCR Assays and Kits for numerous applications including Mutation Detection, Copy Number Determination, Genome Edit Detection, Gene Expression, Expert Design Assays, Residual DNA Quantification and Library Quantification.
-
Image

Featured Product Digital PCR Assays
Get the most out of your experiment with assays designed specifically for Droplet Digital PCR. Simply enter your target of interest or sequence and order a ddPCR Assay.
-
Image

NEW PRODUCT ddPCR Microsatellite Instability (MSI) Kit
The only kit designed to analyze both plasma and FFPE tissue, it provides highly sensitive and reproducible results across all relevant cancer samples without the need for normal tissue. Automated analysis and a streamlined workflow make this the premier MSI solution.
Digital PCR Systems, Software, and Consumables
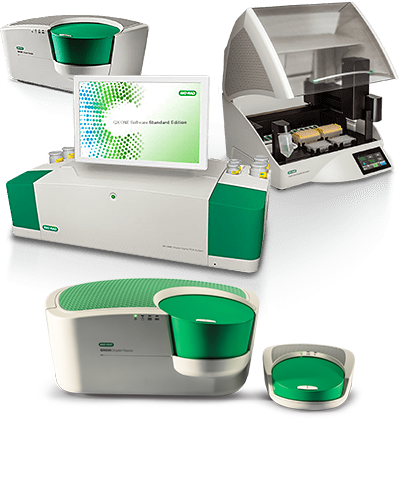

Droplet Digital PCR Systems
Bio-Rad’s Droplet Digital PCR Systems provide ultrasensitive and absolute nucleic acid quantification.
-
QX600 Droplet Digital PCR System
Advanced 6-color multiplexing for up to 96 samples per run
-
QX600 AutoDG Droplet Digital PCR System
Advanced 6-color multiplexing for up to 96 samples per run for high-throughput analysis
-
QX ONE Droplet Digital PCR System
A high throughput multiplexed and integrated droplet digital PCR system
-
QX200 AutoDG Droplet Digital PCR System
For high-throughput analysis
-
QX200 Droplet Digital PCR (ddPCR) System
For analyzing up to 96 samples

-

Droplet Digital PCR System Consumables
Purchase droplet generation oil, cartridges, gaskets, plates & seals, plus pipette tips, and more, designed for Droplet Digital PCR Systems.
Contact a Sales Specialist -
Droplet Digital PCR System Supermix
One-Step RT-ddPCR Advanced Kit for Probes for use with the COVID-19 ddPCR Assay.
Contact a Sales Specialist

QX Software for Digital PCR Systems
QX Software is a family of powerful software packages for acquisition and analysis of your Droplet Digital PCR (ddPCR) data. When connected to a ddPCR System, QX Software provides all necessary functionality to create, run, and analyze experiments on your samples.
Digital PCR Consumables and Reagents
Purchase supermixes, droplet generation oil, cartridges, gaskets, plates & seals, plus pipette tips, and more, designed for Droplet Digital PCR Systems.
-
ddPCR Supermixes
For maximum PCR efficiency and sensitivity. -
ddPCR Oils
For creating sample-in-oil droplets. -
ddPCR Consumables
For use with the droplet generator. -
PX1 Plate Sealer
For secure plate sealing.
QXDx Products for In Vitro Diagnostics (IVD)
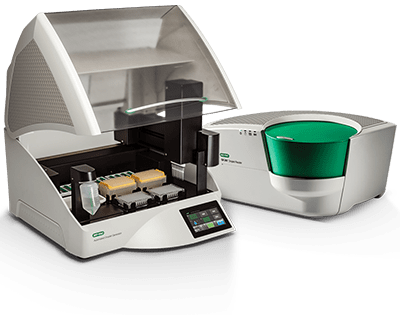
IVD Instrument
CE-IVD Instruments
QXDx AutoDG ddPCR System
- First FDA-cleared ddPCR system for use with QXDx BCR-ABL %IS Kit
- Ultra-low quantification without standard curves
- Droplet partitioning reduces amplification bias
- Automated droplet generator, 96-reaction throughput
- CE-IVD marked ddPCR system for use with the QXDx BCR-ABL %IS Kit
- Ultra-low quantification without standard curves
- Droplet partitioning reduces amplification bias
- Automated droplet generator, 96-reaction throughput
QXDx ddPCR System
- CE-IVD marked ddPCR system for use with the QXDx BCR-ABL %IS Kit
- For analyzing up to 96 samples

IVD Consumables, Assays, and Kits
-

Kits
QXDx Universal Kit for ddPCR System
Reagents and consumables for the IVD ddPCR System
QXDx Universal Kit for AutoDG ddPCR System
Reagents and consumables for the automated IVD ddPCR System
QXDx AutoDG Consumable Pack & QXDx Droplet Reader Oil Pack
Reagents and consumables for the automated IVD ddPCR System.
-

Assays
QXDx BCR-ABL %IS Kit
An FDA-cleared/CE-marked digital PCR test kit intended to monitor patients with Chronic Myeloid Leukemia (CML) with reported results in both International Scale and MR values.
Droplet Digital PCR Technology
Based on water-emulsion droplet technology, Droplet Digital PCR fractionates a DNA sample in 20,000 droplets. PCR amplification of the template subsequently occurs in each individual droplet, and counting the positive droplets gives precise, absolute target quantification.
Read more about Droplet Digital PCR technology or learn how to transition your assay from qPCR to ddPCR.
-

DNA sample with target sequence is partitioned into droplets
-
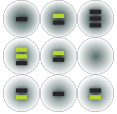

Target and background DNA are randomly distributed among 20,000 droplets
-
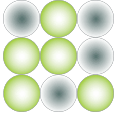

Target sequence is amplified by end-point PCR in each droplet
-
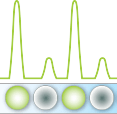
Positive droplets are counted to give precise quantification of target sequences in sample
Benefits of Droplet Digital PCR
Absolute quantification — provides an absolute count of target DNA copies per input sample without the need for standard curves
Highest precision and sensitivity — enriched template concentration in target-positive droplets allows higher precision and sensitivity than real-time PCR
Removal of PCR efficiency bias — error rates are reduced by removing the reliance on amplification efficiency of qPCR
Explore Absolute Quantification with Droplet Digital PCR, download the White Paper on Molecular Counting.
Droplet Digital PCR Applications

Copy number variation (CNV)
Accurate CNV quantification in single wells.
Next generation sequencing
Amplification and accurate quantification of NGS libraries with validation of sequencing results.

Detection of rare sequences in cancer and disease
Detection of mutations and viral load below levels detectable by current tests.

Gene expression analysis
Detection of rare mRNAs and miRNAs with rapid turnover.

Cell & Gene Therapy
ddPCR assays for a range of applications including CAR transgene quantification.

Pathogen detection and microbiome analysis
Viral load and pathogen detection in organ transplants, soil, and food.

Hematology-Oncology
Hematology-oncology solutions provide precise and accurate results without the need for standard curves.

Wastewater-Based Epidemiology
ddPCR assays for unlocking the full potential of wastewater testing and surveillance as a public health tool.
Download the Droplet Digital PCR Applications Guide for in-depth information about setting up experiments.
Bio-Rad is the Trusted Choice for Digital PCR
When it comes to digital PCR, in particular Droplet Digital PCR (ddPCR), Bio-Rad stands as a trusted leader with a track record that speaks for itself. With over 10 years of continuous innovation in digital PCR, and with 8,000+ publications utilizing our ddPCR technology, Bio-Rad has established its reputation as a reliable and respected provider.

8,000+ Publications and Counting
Browse the growing list of cited peer-reviewed journal articles in Droplet Digital PCR research.
Over a Decade of Droplet Digital PCR Innovation and Breakthroughs
-
Image
 2011
2011Bio-Rad Acquires QuantaLife & Digital PCR Technology
This next-generation PCR technology builds on the well-established method of amplifying DNA that is used in research laboratories around the world.
Read the Press Release » -
Image
 2012
2012QX100™ Droplet Digital™ PCR System
Evaluating a treatment for AIDS. Analyzing archival cancer samples. Tracking the RNA of a mutated gene known to cause cancer.
Read the Press Release » -
Image
 2012
2012QX100 Droplet Digital PCR
Bio-Rad releases QX100 Droplet Digital PCR Success Stories video, interviewing technicians from labs using Droplet Digital PCR system.
Watch the Video -
Image
 2013
2013Launch of the QX200™ Droplet Digital™ PCR System
Providing absolute quantification of target DNA or RNA molecules for EvaGreen or probe-based digital PCR applications.
See the System » -
Image
 2013
2013Using Droplet Digital PCR to Diagnose Leukemia
Prof. Alec Morley, a pioneer in digital PCR, describes his use of Droplet Digital PCR (ddPCR) technology to advance the diagnosis and treatment of leukemia.
Watch the Video -
Image
 2014
2014Launch of the QX200™ AutoDG Droplet Digital™ PCR System
The AutoDG Instrument simplifies the Droplet Digital PCR workflow, making digital PCR both scalable and practical.
See the System » -
Image
 2014
2014ddPCR Multiplex Mutation Screening Kit Launch
Designed for rapid screening of several key cancer mutations in a single reaction with high precision and sensitivity.
See the Kit » -
Image
 2014
2014Bio-Rad ddPCR Used in Published Medical Journal Article
A Droplet Digital PCR detection method for rare L1 insertions in tumors.
Read the Article -
Image
 2015
2015Bio-Rad Introduces Droplet Digital™ PCR PrimePCR™ Assay Kits for Spinal Muscular Atrophy Research
This release expands Bio-Rad’s offering of predesigned, fully wet-lab validated assays with kits for SMN1 and SMN2 copy number determination.
Read the Press Release » -
Image
 2015
2015ddPCR Residual DNA Quantification Kits Launch
Bio-Rad’s ddPCR Residual DNA Quantification Kits are ideal for highly precise quantification of HCD in complex bioprocess intermediates.
See the Kits » -
Image
 2015
2015Droplet Digital PCR Proves Highly Reproducible at Identifying Viral RNA Mutations in Clinical Samples
Bio-Rad’s Droplet Digital PCR (ddPCR) technology identifies viral RNA mutations in clinical samples with greater precision, sensitivity, and reproducibility than quantitative reverse-transcription PCR (RT-qPCR), according to a new study in the Journal of Virological Methods.
Read the Press Release » -
Image
 2015
20151-Step RT-ddPCR Advanced Kit for Probes
This digital PCR supermix kit enables absolute quantification of target RNA molecules in a one-step format.
See the Kits » -
Image
 2016
2016The First Digital PCR Instrument with the CE IVD Mark
Bio-Rad announces CE IVD marking of its QX200 Droplet Digital PCR System for use as an in vitro diagnostic.
Read the Press Release » -
Image
 2016
2016Launch of Host Cell Kits & Multiplex Mutation Screening
New application allows precise and accurate cell counts of an edited or modified cell population with minimal input cells.
Read the Bulletin » -
Image
 2016
2016Bio-Rad Launches a Comprehensive Portfolio of Digital PCR Assays and Kits
Applications including Mutation Detection, Copy Number Determination, Genome Edit Detection, Gene Expression, Expert Design Assays, Residual DNA Quantification, and Library Quantification.
See the Kits » -
Image
 2017
2017Bio-Rad Launches QXDx™ Universal Kit for AutoDG ddPCR™ System
Targeted for optimized use with the automated QXDx System, this kit enables users to complete every step of the automated Droplet Digital PCR in vitro diagnostics workflow with high efficiency.
See the Kit » -
Image
 2017
2017Bio-Rad Unveils QXDx BCR-ABL %IS Kit (IVD/CE-IVD)
Intended to monitor patients with Chronic Myeloid Leukemia (CML), this FDA-cleared ddPCR kit reports in both International Scale (%IS) and Molecular Response (MR).
See the Kit » -
Image
 2017
2017Bio-Rad Releases Paper on ddPCR Genome Edit Detection Assays
Assays offer a fast, precise, simple, and cost-effective method for detection of genome editing events.
Read the Bulletin » -
Image
 2019
2019The QXDx AutoDG ddPCR System is Shown for the First Time
Designed for high-throughput processing of samples in a 96-well format, this system simplifies the ddPCR workflow and eliminates user-to-user variability in IVD applications.
See the System » -
Image
 2019
2019Bio-Rad Releases First FDA-Cleared Digital PCR System & Test for Monitoring Chronic Myeloid Leukemia Treatment Response
The QXDx AutoDG ddPCR System and QXDx BCR-ABL %IS Kit represent the first-ever digital PCR solution that can monitor and directly quantitate the molecular response of patients with chronic myeloid leukemia under tyrosine kinase inhibitor therapy.
Read the Press Release » -
Image
 2020
2020QX ONE™ Droplet Digital™ PCR (ddPCR™) System Release
A high throughput, multiplexed and integrated Droplet Digital PCR Sytem.
See the System » -
Image
 2020
2020Bio-Rad Receives FDA Emergency Use Authorization for Droplet Digital PCR SARS-CoV-2 Test Kit
The SARS-CoV-2 Droplet Digital PCR (ddPCR) test runs on Bio-Rad's QX200 and QXDx ddPCR Systems.
Read the Press Release » -
Image
 2021
2021Vericheck ddPCR Mycoplasma Detection Kit Release
The probe-based Vericheck ddPCR Mycoplasma detection Kit brings confidence in testing for mycoplasma contamination. The specificity and reproducibility of this kit allows for a precise measurement of 112 Mycoplasma species using Droplet Digital PCR.
See the Kit » -
Image
 2022
2022Launch of the QX600™ Droplet Digital™ PCR System
The QX600 System provides advanced, 6-color multiplexing to maximize the number of biomarkers interrogated per well, allowing the preservation of precious samples and reducing lab burden.
See the System »
- 2011
- 2012
- 2012
- 2013
- 2013
- 2014
- 2014
- 2014
- 2015
- 2015
- 2015
- 2015
- 2016
- 2016
- 2016
- 2017
- 2017
- 2017
- 2019
- 2019
- 2020
- 2020
- 2021
- 2022

Empower Your Research with ddPCR Technology
Our commitment to advancing the field of molecular diagnostics is reflected in the accuracy, sensitivity, and reliability of our ddPCR solutions, making Bio-Rad the trusted choice for researchers worldwide. Join the growing community of scientists who rely on Bio-Rad's ddPCR technology for their groundbreaking discoveries.
Contact a Sales Specialist


























