On This Page |
What is Stain-Free |
Advantages of Stain-Free |
Total Protein Normalization |
FAQs | Resources | Related Products |
Consult with a Specialist |
What is Stain-Free Imaging?
Stain-Free imaging technology utilizes a polyacrylamide gel containing a proprietary trihalo compound to make proteins fluorescent directly in the gel with a short photoactivation, allowing the immediate visualization of proteins at any point during electrophoresis and western blotting. This trihalo compound is covalently bound to tryptophan residues, enhancing their fluorescence when exposed to UV light, enabling the detection of proteins at levels as low as 10–25 ng.
The addition of the fluorophore allows visualization of proteins in the gel and following transfer onto a membrane during western blotting but does not interfere with electrophoresis or downstream steps. With the fluorophore covalently bound to the protein molecules, they can be imaged repeatedly on a gel or membrane after transfer, without additional staining and destaining steps.
Stain-Free imaging allows for the elimination of the inherently problematic use of housekeeping proteins as loading controls on western blots, enabling the user to obtain truly quantitative western blot data by normalizing bands to total protein in each lane.
Making a Stain-Free Gel

Making a Stain-Free Gel
A trihalo compound is mixed with acrylamide solution before casting the gel. Alternatively, commercially available Stain-Free Gels can be used. These incorporate the trihalo compound in their gel chemistry or are provided as gel solutions for hand casting.
Total protein visualization on gel
The trihalo compound modifies tryptophan residues in the protein by a covalent modification, generating a fluorescence signal. The signal is visualized by UV excitation. The trihalo compound by itself does not produce any fluorescence and so no background signal is obtained.
Visualization without further activation on blot
Nitrocellulose or PVDF membranes generate autofluorescence. Low-fluorescence PVDF can be used to reduce background.
Advantages of Using Stain-Free Technology
Stain-Free Technology Provides More Sensitivity and Better Dynamic Range than Coomassie Stains
Protein visualization data obtained from Stain-Free gels are comparable to those obtained from gels stained with other dyes. In general, the sensitivity of Stain-Free gels when visualizing data is equal to that of Coomassie-stained gels for all proteins. For proteins with higher tryptophan content, Stain-Free gels provide much higher sensitivity than CBB-stained gels (Figure 1).

Figure 1. Comparison of a Stain-Free gel, CBB R-250, and Bio-Safe G-250 stained gel images. Serial 1:2 dilutions of broad range unstained molecular weight standards were separated on a 4–20% Criterion Stain Free Tris-HCI Gel. The gel was imaged with a Stain-Free enabled imager, then stained with Coomassie (CBB R-250 and Bio-Safe G-250) stain and imaged on a densitometer. Arrowhead indicates β-galactosidase.
The limit of detection for the Stain-Free gels is 8 to 28 ng, similar to that of silver stains (0.6 to 1.2 ng), while Coomassie R-250 stain can detect protein amounts of at least 35 to 50 ng. Some fluorescent stains can detect proteins at levels below the 1 or 0.5 ng limit. Stain-Free gels have more reproducible data with smaller coefficients of variation compared to Coomassie or silver stains (McDonald et al. 2008; McDonald, 2009).
The linear dynamic range for protein quantitation is defined as the range through which the signal intensity on a blot proportionally increases with the increase in protein load. Ideally, the protein load should fall within the quantitative linear dynamic range of the antibody used for its detection (Taylor and Posch, 2014). Stain-Free gels provide a linear dynamic range between 10 and 80 µg of total protein load from cell or tissue lysates at a higher range of protein load (Figure 2A) and from 20 to 1 µg at a lower range (Figure 2B) (Taylor et al. 2013, Hammond et al. 2013).

Figure 2. Linear dynamic range provided by Stain-Free technology for total protein measurements. A. HeLa cell lysate dilutions from 80–2.5 µg total protein; B. HeLa cell lysate dilutions from 20–1 µg total protein.
Compatibility with Downstream Applications
The Coomassie dye-based method does not allow the same gel to be used for transfer during western blotting or for mass spectrometry. This is a serious drawback when the ultimate goal is the identification or quantitation of proteins in western blotting, especially when large gels and sample volumes are required for separation and visualization. In contrast to Coomassie staining, Stain-Free technology is compatible with most downstream applications, such as western blotting, which helps not only when resources are limited, but also in the appropriate normalization of data for quantitation using total protein normalization (see Normalization of Data in Quantitative Western Blotting below).
Verification of Protein Transfer
The proper transfer of proteins to the membrane is critical to the western blotting process. A common standard practice to verify protein transfer is staining the blot with Ponceau S, a negatively charged stain that binds to all positively charged amino acids in a protein. Ponceau S staining is fast and relatively inexpensive, and the stained membrane can then be used for downstream applications such as western blotting. However, the ephemeral nature of the binding of the dye to the protein causes the intensity of the bands on the membrane to decrease rapidly, making detection and quantitation harder. Other relatively expensive blot stains, such as SYPRO Ruby or amido black, are also used for blot quantitation and require special disposal methods. The time needed to view the stain on the blot also varies with the dye used. Protocols for using Ponceau stain recommend at least 5 minutes of staining followed by 15 minutes of washing steps. SYPRO Ruby Blot Stain is more elaborate, requiring fixation, overnight staining, and destaining.
Verifying protein transfer, both from the gel and on the membrane, using a Stain-Free enabled imager provides significant time savings and cost savings for western blotting (Colella et al. 2012). The observed intensity of the bands does not depend on the duration of staining or destaining, a factor affecting dye-based techniques during visualization and quantitation. Also, the intensity of the band on Stain-Free blots, which is the result of a covalent modification, does not decrease with time. Using the Stain-Free method, protein transfer can be verified in as little as 2 minutes.
-
Gel before transfer
-

Membrane after transfer
-

Gel after transfer
Assessment of protein transfer using a Stain-Free enabled imaging system. Images of the gel before and after transfer and of the membrane after transfer were taken using the Criterion Stain-Free Imager. Serial 1:2 dilutions of hemoglobin (starting quantity, 80 ng), with 1.8 μg of BSA/lane as a carrier (top band), were electrophoretically separated on a 4–20%, 26-well Criterion Stain-Free Gel.
Total Protein Normalization
Using Stain-Free total protein measurement as the loading control, researchers can ensure that both the target protein and loading control are measured in the linear dynamic range in a typical western blot experiment. Furthermore, Stain-Free imaging allows for the complete elimination of the inherently problematic use of housekeeping proteins as loading controls on western blots. Stain-Free total protein measurement serves as a more reliable loading control than housekeeping proteins, particularly in the loading range commonly used for cell lysates, permitting the user to obtain truly quantitative western blot data by normalizing bands to total protein in each lane.
Normalization across a wider dynamic range
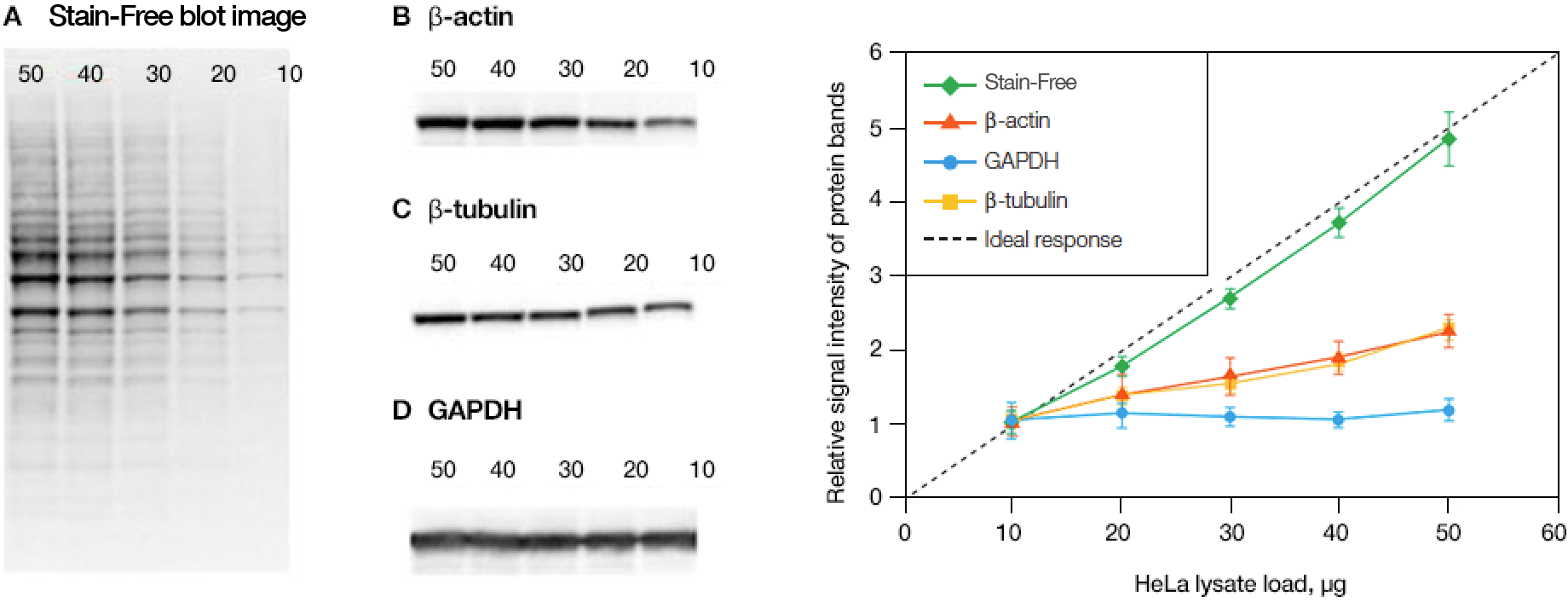
Linearity comparison of Stain-Free total protein measurement and immunodetection of three housekeeping proteins in 10–50 μg of HeLa cell lysate. On the left are representative images of (a) Stain-Free blot and the chemi blots for (b) β-actin, (c) β-tubulin, and (d) GAPDH. Lane labels correspond to total protein load (μg). Although the actin and tubulin signals appear linear, the densitometric ratio was far below the predicted “quantitative response” of actual loading whereas the Stain-Free signal correlated to the expected result (e).
Taylor SC Posch A (2014). The design of a quantitative western blot experiment. Biomed Res Int. 36, 1590.
Advantages of Total Protein Normalization
Stain-Free total protein normalization is performed by measuring total protein directly on the membrane that is used for western blotting. The total density for each lane is measured from the blot and a lane profile is obtained. The background is adjusted in such a way that the total background is subtracted from the sum of density of all the bands in each lane (referred to as the rolling disk background subtraction algorithm).
Total protein normalization takes into account:
- The intensity of all proteins in the lane
- Sample loading variations in each lane
- Experimental variations during electrophoresis, transfer and blotting
Stain-Free Imaging of Chemiluminescence and Multiplex Fluorescence Western Blots
Stain-Free gels are fully compatible with both chemiluminescence and fluorescent western blotting detection methods. For chemiluminescence detection, acquire blot image on Stain-Free enabled imager just before applying ECL reagent to achieve the most accurate representation of total protein present.
-

Stain-Free Image of Blot
-

Chemiluminescence Image of Blot
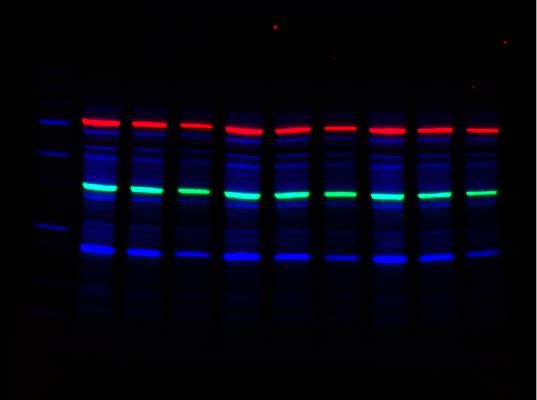
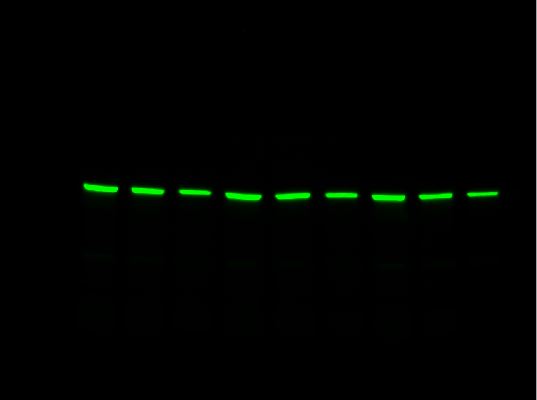
For multiplex western blot applications, use the red, far red, and near-IR channels for detecting proteins of interest and reserve the blue channel for total protein detection using Stain-Free technology.
-
Overlay
-
Stain-Free Channel
-
Red Channel
-
Far-Red Channel
FAQs
Q: Will I need to purchase new buffers and reagents to utilize Stain-Free technology?
A: Stain-Free gels do not use specialized buffers or reagents. Standard SDS-PAGE buffers can be used. However, the technique does require an imager that can visualize the activated gel or membrane. View our Stain-Free Imaging Systems.
Q: As Stain-Free technology utilizes the modification of tryptophan residues, will it work on my protein of interest?
A: Theoretically, even one tryptophan residue is sufficient for signal activation, and proteins that have as few as two tryptophan residues are readily detected and quantified using Stain-Free technology. For most applications, the requirement of tryptophan residues for visualization is not a concern. Data available from UniProt show that only about 10% of proteins from all organisms lack tryptophan and most of those proteins are less than 10 kD in size. In most organisms, approximately 90% of proteins are above 10 kD in size (in the 10–260 kD range) and possess tryptophan residues (Table 1).
Table 1. Tryptophan content of the predicted proteomes of several model organisms.*
| Species | Total Number of Proteins | Number of Proteins Lacking Tryptophan | % of Proteins Lacking Tryptophan | Number of Proteins >10 kD | Number of Proteins > 10 kD Lacking Tryptophan | % of Proteins >10 kD Lacking Tryptophan |
| Homo sapiens | 40,827 | 4,209 | 10.31 | 37,548 | 2,754 | 7.33 |
| Escherichia coli O:K1 / APEC | 4,865 | 458 | 9.41 | 4,754 | 408 | 8.58 |
| Escherichia coli (strain K12) | 4,181 | 456 | 10.91 | 3,879 | 325 | 8.38 |
| Rattus norvegicus | 12,022 | 1,081 | 8.99 | 11,421 | 745 | 6.52 |
| Mus musculus | 35,344 | 3,435 | 9.72 | 33,262 | 2,480 | 7.46 |
| Saccharomyces cerevisiae | 5,815 | 648 | 11.14 | 5,563 | 491 | 8.83 |
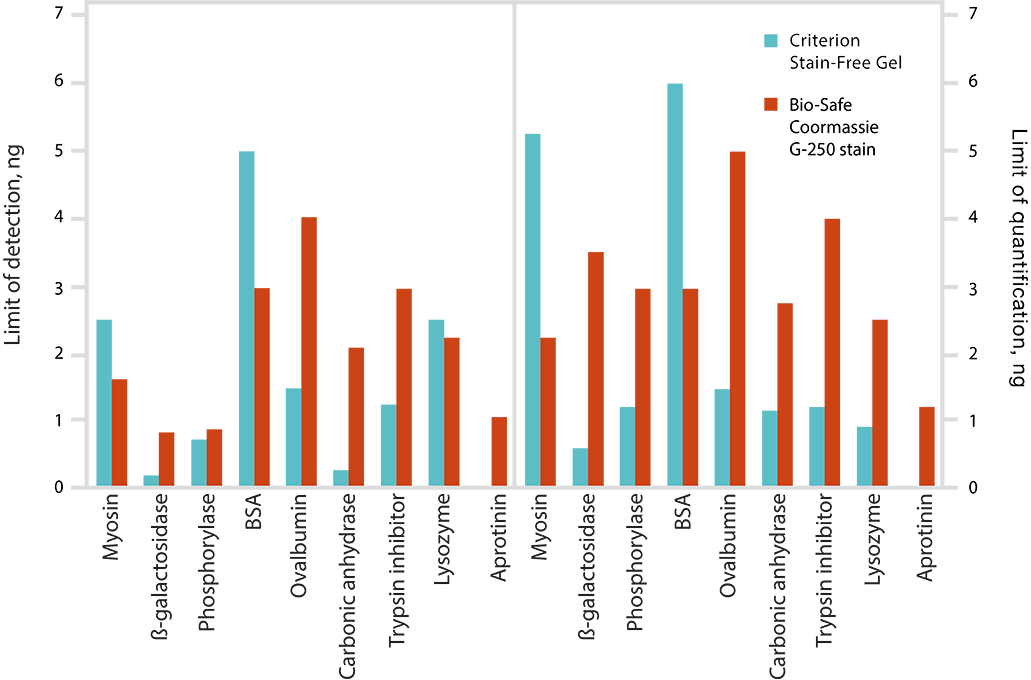
Limits of detection (LOD) and limits of quantitation (LOQ) of proteins on Criterion Stain-Free Gels and Bio-Safe G-250 Stained Gels. Individual bands from broad range unstained protein standards from four replicate gels were used to determine visual LOD and LOQ. Averaged numbers were used to generate the graph.
Q: How does Stain-Free affect the way we do protein research today?
A: With an increasing number of errors in quantitation arising due to the use of housekeeping proteins and associated normalization issues, a growing number of journals in the western blotting field have forced authors to withdraw published reports, issue errata, or have their publications rejected (Neill 2009). Journals are now requiring strict adherence to the use of internal controls and are mandating the use of imaging techniques that yield linear signal ranges and report linear dynamic range of signal (see guidelines for reporting life sciences research from Nature Publication Group 2013 and guidelines from Journal of Biological Chemistry). Stain-Free technology fits perfectly in this new world of western blotting as it satisfies all the requirements for carrying out these new protocols and is considered an effective strategy to improve accuracy of western blots (Ghosh et al. 2014).
Meeting the New Publication Requirements for Quantitative Western Blotting
| JBC Guidelines: Quantitative Blots | How to Meet Requirements | Bio-Rad Western Workflow Solutions |
|---|---|---|
| "Housekeeping proteins should not be used for normalization without evidence that experimental manipulations do not affect their expression" | Use Total Protein Normalization (TPN) instead of housekeeping proteins | Stain-Free Western Blotting simplifies TPN. This technology is proven with over 100 publications, allows for improved accuracy and will make your data more reliable |
| "Methods including detection of enhanced chemiluminescence using X-ray film have a very limited dynamic range" | Use an imaging system with at least 4 logs of dynamic range | ChemiDoc Systems are Stain-Free imagers with the sensitivity of film Image Lab Software enables simple image generation and quantitation |
| "A description of the data supporting the specificity of all antibodies is required" | Use fully validated antibodies | PrecisionAb Antibodies offer documented specificity and validation protocols |
Sign Up to Be the First to Know
Receive Western Blotting Tips and Online Resource Updates
Keep up to date with useful tips to continuously improve your western blotting experiments from sample preparation through image analysis. Sign up to be the first to be notified when new western blotting resources like tips and tricks, posters, protocols, webinars, and how-to-videos become available.
Resources

Free eBook
Find out how Stain-Free technology can revolutionize your western blotting
Request Yours Now
Total Protein Normalization
Learn how total protein normalization is key to obtaining quantitative western blot data for publication.

Fluorescent Western Blotting
See how fluorescent western blotting addresses the need for accurate, quantitative determination of protein expression.

Film vs. Digital Western Blot Imaging
Discover how modern digital imaging systems, with wide dynamic range enable, more accurate quantitation over a range of signal intensities that are equal to or better than film.

Image Lab Touch Software Resources
Access Image Lab Touch Software resources such as: instructional videos, information on Smart Tray Technology, and tips on capturing images including exporting data.
Image Lab Software Resources
Bio-Rad provides tutorials for basic acquisition and advanced analysis use of Image Lab Software 6.0.1. Topics include, densitometric analysis, molecular weight, normalization, purity assessment, and more!

Crucial Factors of Successful Western Blotting
Join our western blotting expert and guest speakers for a webinar series exploring factors that go into the design and execution of successful, repeatable western blots. Chemiluminescence or fluorescence, qualitative or quantitative blot — we’ll discuss how-to's and take a deep dive exploring the why's.
Documents Library

ChemiDoc MP System Brochure
(PDF 1.26 MB)Explore the features and benefits of the new high-end imaging system for the best fluorescence and chemiluminescence detection.

Stain-Free Approach to Western Blotting
(PDF 864 KB)Learn how to use Stain-Free technology for total protein normalization as an alternative to the standard blot normalization process.

Stain-Free Western Workflow Brochure
(PDF 593 KB)See how this improved workflow incorporating Stain-Free gel technology saves time and increases the accuracy and reliability of your western blot results.

Determining the Appropriate Sample Load When Using a Stain-Free V3 Western Workflow
(PDF 182 KB)Determine the right load for the detection of target proteins before the actual western blot experiment is carried out.

Validating the Expression Consistency of a Housekeeping Protein
(PDF 198 KB)Validate the consistency of HPK expression before the actual western blot experiment is performed.
Related Products

Stain-Free Western Blotting
A streamlined approach to conventional western blotting; incorporating Stain-Free TGX gels enabling faster results, better data.

ChemiDoc Imaging Systems
ChemiDoc Imagers offer best-in-class performance for fluorescence and chemiluminescence detection and all general gel documentation applications.
Image Lab Software
Image acquisition and analysis software for Bio-Rad Gel Doc, ChemiDoc, and GS-900 Systems. Capture and analyze digital image data from electrophoresis gels and blots.
Western Blotting
Western blotting products include Stain-Free Western Blotting, protein transfer systems, membranes, filter paper, blotting buffers, and detection kits.
Fluorescent Western Blotting
Bio-Rad's fluorescent western blotting workflow is a seamless integration of products designed to work together to offer guaranteed results.

Trans-Blot Turbo Transfer System
The Trans-Blot Turbo System is a rapid protein transfer apparatus that can transfer protein to membrane in as little as 3 minutes. Trans-Blot Turbo Transfer Packs provide greater transfer efficiency in less time.
PrecisionAb Validated Western Blotting Antibodies
These premium antibodies are lab-validated using strict testing criteria to ensure superior performance in western blotting detection.

Mini-PROTEAN TGX Stain-Free Precast Gels
Mini-PROTEAN TGX Stain-Free Precast Gels for polyacrylamide gel electrophoresis (PAGE) allow for fast run times, high transfer efficiency, and rapid detection of proteins with Stain-Free enabled imagers.
Protein Ladders and Standards (Markers)
Prestained and unstained molecular weight standards for protein electrophoresis applications including SDS-PAGE, western blotting, 2-D PAGE, and isoelectric focusing (IEF).

Clarity ECL Substrates
Clarity family of Western ECL Substrates provides simple, high-performance solutions for all your western chemiluminescence needs.