Listed below are some common symptoms in an allelic discrimination experiment. Select one of the symptoms to view possible causes and solutions.
Related Topics: Gene Expression/Quantification Experiments and High Resolution Melt (HRM) Experiments.
More Than Three Clusters
Back to TopDisplay Mode
Back to Top|
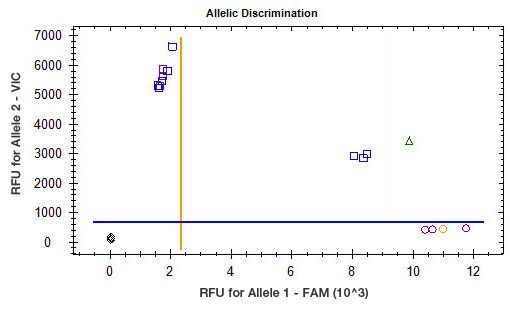
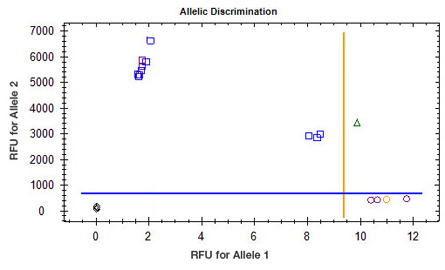
CFX Maestro Software has two different display modes, quantification cycle (Cq) and relative fluorescence units (RFU). The Cq display mode displays the data as Cq values based on the selected thresholds; the RFU display mode displays the data in relative fluorescence units at the selected cycle. |
|
| Possible Solutions | Try both display modes to determine the best mode for your data set. |
Threshold
Back to Top|
Depending on your data set, the threshold may need to be adjusted. CFX Manager software automatically sets the threshold lines for discriminating alleles by taking an average Cq or RFU from your positive controls. If the run contains three controls in the plate, then the position of the threshold bars is based on the mean and standard deviation of the RFU or Cq of the controls. However, if the number of controls is less than three, then the position of the threshold bars is determined by the range of RFU or threshold cycle values in the selected fluorophore. |
|
| Possible Solutions | Click and drag the threshold bars in the Allelic Discrimination chart to adjust the thresholds. |
| Manually adjust the Cq threshold in the amplification chart. | |
Positive Controls
Back to Top|
Some samples may be difficult to discriminate without positive controls. Ensuring that you have a positive control for each allele will help set the thresholds to discriminate between the different alleles. |
|
| Possible Solutions | Add more positive controls. |
Homozygotes as Heterozygotes
Back to TopInaccurate calling of homozygotes as heterozygotes may be due to the causes below. Select one of the causes for possible solutions.
Contamination
Back to Top|
The ability to detect and amplify very small numbers of starting templates by PCR means that contamination control is always important. Contamination can cause reactions that should otherwise fail to be positive for the allele. Contamination in master mix components will show up in all samples. Contamination during plate loading typically affects one or a few wells. |
|
| Possible Solutions | Include no-template control (NTC) samples to help detect global contamination, such as contamination of master mix components. |
| Set up work space to avoid contamination. Do not work with contamination sources, such as post-PCR products, in the same area where PCR setup is done. | |
| Run replicate samples (duplicates, triplicates, etc.) to help identify when loading contamination may have occurred. | |
| Load plate carefully and do not try to expel the residual volume from the pipet tip because this can cause bubbles which may spray droplets into adjacent wells. | |
| Use barrier tips to avoid contamination due to your pipet. | |
Long Amplicons
Back to Top|
The length of an amplicon can influence the amplification and melt profile especially if the sequence contains more than one mutation. As a result, the mutation of interest can be difficult to separate from the other sequence variants if primer sets are designed for longer amplicons (300 bp). |
|
| Possible Solutions | Use amplicons with a target length <300 bp to improve detection of sequence variants. |
Nonspecific Amplification
Back to Top|
Allelic discrimination requires well-optimized assays. Sometimes nonspecific amplification can occur, which may lead to false positive signals for one or more alleles. Nonspecific amplification can occur due to the loss of hot-start enzyme activity, degraded reaction components, or too much starting template. |
|
| Possible Solutions | Use amplicons with a target length <300 bp to improve detection of sequence variants. |
| Include NTC samples to detect template-independent reactions (for example, primer/probe dimers) and reoptimize assays if problems are detected. | |
| Optimize assay conditions using real samples whenever possible. Cross- reactivity to nontarget alleles (for example, pseudogenes) can be problematic for assays developed using cloned products or other simplified targets. | |
| Prior to hot-start activation, use a qPCR supermix with good inhibition of polymerase activity. | |
| Store the PCR components properly; especially take care to aliquot the probes in sufficiently small volumes to avoid multiple freeze-thaw cycles, and store the aliquots below –20°C. | |
| Try to keep the amount of starting template similar among reactions. | |
Incorrect Positioning of Thresholds
Back to Top|
The automatic calling of alleles is determined by the positioning of the threshold lines. It is therefore essential that these thresholds be set appropriately for the alleles to be called correctly. CFX Manager software typically positions the threshold automatically, but it may need to be adjusted in some cases. The autothresholds are most likely to be placed incorrectly when the variation among unknown samples differs from that among control samples. The Allelic Discrimination tab allows the data to be displayed as Cq values or RFU values, depending on your preference. Note that positive samples typically have lower Cq values and higher RFU values than negative samples. |
|
| Possible Solutions | Include the appropriate control samples. It is helpful to include not only positive control samples for each allele, but also NTCs and heterozygotes. Control samples should be treated as similarly to the unknowns as possible. |
| Adjust the threshold lines manually by clicking and dragging when necessary. To distinguish between a true positive signal and a background signal it can often be helpful to view the amplification plots on the Quantification tab. Thresholds must be set correctly in the Quantification tab to achieve proper Cq values. Note that the thresholds on the Allelic Discrimination tab are independent of the thresholds on the Quantification tab. Thresholds on the Quantification tab only affect the Cq values, not RFU values. | |
| View the data in both Cq and RFU modes to see which allele calls change. Allele calls that change typically warrant closer examination of the underlying data or manually adjusting thresholds. | |
| Consider well-to-well differences (such as amplification efficiency and starting template amounts) when setting thresholds to better account for variation between samples, especially if the control samples were extracted or otherwise treated differently. Thresholds should fall between all the positive and negative controls for that allele, but only rarely midway between them. | |
Control Samples Are Not Homozygotes
Back to Top|
It is important to use control samples with known genotypes. A variety of causes can lead to improperly identified control samples, including mislabeling, misidentifying the sample or the genotype, overgrowth or contamination of cultured material, and mutation. |
|
| Possible Solutions | Verify genotypes of source material. |
| Rerun reactions to identify if use of incorrect material or incorrect labeling occurred previously; pseudogenes can be problematic for assays developed using cloned products or other simplified targets. | |
Further Reading

Castellanos E et al. (2010). Rapid identification and differentiation of Mycobacterium avium subspecies paratuberculosis types by use of real-time PCR and high-resolution melt analysis of the MAP1506 locus. J Clin Microbiol 48, 1474–1477. PMID: 20129970
Gan XL et al. (2010). Association of an interleukin-16 gene polymorphism with the risk and pain phenotype of endometriosis. DNA Cell Biol 29, 663–667. PMID: 20662556