Proteins separated by gel electrophoresis can be visualized using different staining procedures. The choice of staining technique depends on the availability of imaging equipment in the lab in many cases. This section provides general considerations for staining and describes different total protein stains such as Coomassie, fluorescent, silver, and negative stains, and provides troubleshooting tips and protocols for different staining procedures. It also discusses the features and benefits of stain-free technology for protein visualization.
Related Topics: SDS-PAGE Analysis.
Page Contents
A protein staining technique should offer the following:
- High sensitivity and reproducibility
- Wide linear dynamic range
- Compatibility with downstream technologies such as protein extraction and assay, blotting, or mass spectrometry
- Robust, fast, and uncomplicated protocol
Staining protocols usually involve the following three steps:
- Protein fixation, usually in acidic methanol or ethanol (a few staining protocols already contain acid or alcohols for protein fixation and do not require this separate step)
- Exposure to dye solution
- Washing to remove excess dye (destaining)
See the protocols tab below for individual protocols.
Total protein stains allow visualization of the protein pattern in the gel. The table below compares the advantages and disadvantages of several total protein staining techniques.
Coomassie Stains
The most popular anionic protein dye, Coomassie Brilliant Blue, stains almost all proteins with good quantitative linearity at medium sensitivity. There are two variants of Coomassie Brilliant Blue: R-250 (R for reddish), which offers shorter staining times, and G-250 (G for greenish), which is available in more sensitive and environmentally friendly formulations (Simpson 2010, Neuhoff et al. 1988). Coomassie dyes are also the favorite stains for mass spectrometry and protein identification. Bio-Safe Coomassie Stain is a nonhazardous formulation of Coomassie Blue G-250 that requires only water for rinsing and destaining. It offers a sensitivity that is better than conventional Coomassie R-250 formulations and equivalent to Coomassie Blue G-250, but with a simpler and quicker staining protocol.
Silver Stains
Silver stains offer the highest sensitivity, but with a low linear dynamic range (Merril et al. 1981, Rabilloud et al. 1994). Often, these protocols are time-consuming, complex, and do not offer sufficient reproducibility for quantitative analysis. In addition, their compatibility with mass spectrometry for protein identification purposes is lower compared to Coomassie stains and fluorescent dyes (Yan et al. 2000). Silver stain offers sensitivity that exceeds that of Coomassie and is equivalent to most fluorescent stains.
Fluorescent Stains
These stains fulfill almost all the requirements for an ideal protein stain by offering high sensitivity, a wide linear dynamic range over four orders of magnitude, a simple and robust protocol, and compatibility with mass spectrometry. In comparison to Coomassie or silver staining techniques, however, fluorescent dyes are more expensive and require either a CCD (charge-coupled device) camera or fluorescence scanner for gel imaging. For these reasons, fluorescent stains are often used in proteomics applications and on 2-D gels, where the relative quantitation of proteins in complex mixtures over several orders of abundance is performed and protein identity is determined using in-gel proteolytic digestion and mass spectrometry. Examples include Flamingo™ and Oriole™ Fluorescent Gel Stains.
Stain-free technology enables the visualization of proteins in gels and on membranes without a staining step and provides several advantages over conventional staining procedures. In particular, it eliminates the need for a lengthy destaining step, which makes visualization faster and more efficient and eliminates the need to dispose of organic waste generated during destaining.
Stain-free technology also offers tremendous advantages in quantitative western blotting experiments. Using a stain-free enabled imager, total protein normalization can be performed on posttransfer membranes, allowing for precise quantitation of western blotting signals. Total protein normalization is becoming the gold standard for western blotting normalization in major peer-reviewed journals.
Learn more about the advantages of using stain-free technology.
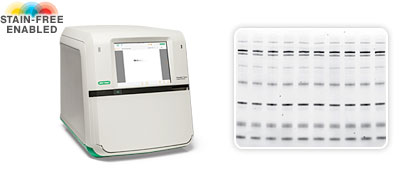
Bio-Rad's ChemiDoc Touch Stain-Free Enabled Imaging System. Left: the ChemiDoc Touch System comprises the imager with UV/stain-free tray and Image Lab Software. Right: Protein gel imaged using stain-free technology.
Specific protein stains are used to visualize specific protein classes such as glycoproteins (Hart et al. 2003) and phosphoproteins (Steinberg et al. 2003, Agrawal and Thelen 2009), which are of special interest to researchers working in the life sciences (examples include Pro-Q Diamond and Pro-Q Emerald).
Bio-Rad Gel Stain Selection Guide

| Total Protein Stain | Sensitivity (Lower Limit) |
Time | Comments | Imaging | |
| Stain-Free | |||||
| Similar to Coomassie and dependent upon tryptophan | No staining or N/A | Fast and reproducible; visualize proteins in 5 minutes or less with a stain-free enabled imager; no staining required | |||
| Coomassie Stains | Densitometer | ||||
| Bio-Safe Coomassie G-250 | 8–28 ng | 1–2.5 hr | Nonhazardous staining in aqueous solution; premixed, mass spectrometry compatible | ||
| Coomassie Brilliant Blue R-250 | 36–47 ng | 2.5 hr | Simple and consistent; mass spectrometry compatible; requires destaining with methanol | ||
| QC Colloidal Coomassie | 3 ng | 1–20 hr | Colloidal endpoint stain; premixed, nonhazardous formulation — no methanol required | ||
| Silver Stains | Densitometer | ||||
| Silver Stain Plus™ Kit | 0.6–1.2 ng | 1.5 hr | Simple, robust; mass spectrometry compatible (Gottlieb and Chavko 1987) | ||
| Fluorescent Stains | CCD and Laser-Based Scanners | ||||
| Oriole Fluorescent Gel Stain* | 0.5–1 ng | 1.5 hr | Rapid protocol, requires no destaining, mass spectrometry compatible; compatible only with UV excitation | ||
| Flamingo Fluorescent Gel Stain | 0.25–0.5 ng | 5 hr | High sensitivity; broad dynamic range; simple protocol requires no destaining; mass spectrometry compatible; excellent for laser-based scanners | ||
| SYPRO Ruby Protein Gel Stain | 1–10 ng | 3 hr | Fluorescent protein stain; simple, robust protocol; broad dynamic range; mass spectrometry compatible | ||
* Do not use Oriole gel stain with native gels.
Staining Equipment
Commercially available stainers such as Dodeca™ High-Throughput Stainers can be used in staining procedures to increase consistency in staining and to avoid breakage of gels from excess handling.

Dodeca High-Throughput Stainers.
| Problem | Cause | Solution |
| Bands not visible | No protein in gel | Stain with another method to confirm there is protein |
| Imaging system malfunctioning | Check instrument manual for troubleshooting, or contact imaging instrument manufacturer | |
| Incorrect imaging parameters were used | Check instrument manual | |
| Poor staining sensitivity | Dirty staining trays | Clean staining trays and other equipment with laboratory glassware cleaner |
| Insufficient stain volume | Follow recommendations for stain volume (appropriate to gel size) | |
| Insufficient staining time | Increase staining time | |
| Reuse of staining solution | Repeat staining protocol with fresh staining solution | |
| High or uneven background staining | Dirty equipment or staining trays | Clean staining trays and other staining equipment with laboratory glassware cleaner |
| Too much time in staining solution | Restrict duration of incubation in staining solutions as recommended in protocol Wash gel in water or respective destaining solution for ≥30 min |
|
| Reagent impurities | Use high-purity water and reagents for staining | |
| Speckles or blotches in gel images | Particulate material from reagents, staining tray, dust or gloves | Clean staining trays thoroughly Limit time that gels and staining solution are exposed to open air Use dust-free gloves and handle gels only by edges |
| Uneven staining | Insufficient shaking during staining | Agitate gel during staining |
| Gel shrinkage | Transfer gel to water for rehydration |
Agrawal GK and Thelen JJ (2009). A high-resolution two dimensional Gel- and Pro-Q DPS-based proteomics workflow for phosphoprotein identification and quantitative profiling. Methods Mol Biol 527, 3–19.
Gottlieb M and Chavko M (1987). Silver staining of native and denatured eukaryotic DNA in agarose gels. Anal Biochem 165(1), 33–37.
Hart C et al. (2003). Detection of glycoproteins in polyacrylamide gels and on electroblots using Pro-Q Emerald 488 dye, a fluorescent periodate Schiff-base stain. Electrophoresis 24, 588–598.
Merril CR (1987). Detection of proteins separated by electrophoresis. Adv Electrophoresis 1, 111–139.
Merril CR et al. (1981). Ultrasensitive stain for proteins in polyacrylamide gels shows regional variation in cerebrospinal fluid proteins. Science 211, 1437–1438.
Miller I et al. (2006). Protein stains for proteomic applications: which, when, why? Proteomics 6, 5385–5408.
Neuhoff V et al. (1988). Improved staining of proteins in polyacrylamide gels including isoelectric focusing gels with clear backgrounds at nanogram sensitivity using G-250 and R–250. Electrophoresis 9, 255–262.
Rabilloud T et al. (1994). Silver-staining of proteins in polyacrylamide gels: a general overview. Cell Mol Biol 40, 57–75.
Simpson RJ (2010). Rapid coomassie blue staining of protein gels. Cold Spring Harb Protoc, pdb prot5413.
Steinberg TH et al. (2003). Global quantitative phosphoprotein analysis using multiplexed proteomics technology. Proteomics 3, 1128–1144.
Yan JX et al. (2000). A modified silver staining protocol for visualization of proteins compatible with matrix-assisted laser desorption/ionization and electrospray ionization-mass spectrometry. Electrophoresis 21, 3666–3672.
Documents
TEST
| 6209 | Total Protein Staining | Click to download |
| 6217 | Total Protein Detection | Click to download |
